Structural insights into H(+)-coupled multidrug extrusion by a MATE transporter
Lu, M., Radchenko, M., Symersky, J., Nie, R., Guo, Y.(2013) Nat Struct Mol Biol 20: 1310-1317
- PubMed: 24141706 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2687
- Primary Citation Related Structures:
4LZ6, 4LZ9 - PubMed Abstract:
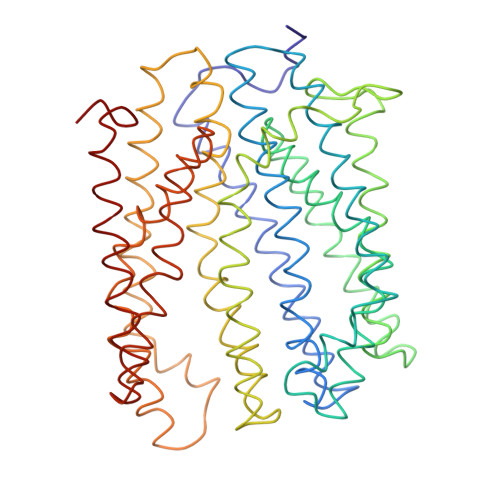
Multidrug and toxic compound extrusion (MATE) transporters contribute to multidrug resistance by coupling the efflux of drugs to the influx of Na(+) or H(+). Known structures of Na(+)-coupled, extracellular-facing MATE transporters from the NorM subfamily revealed 12 membrane-spanning segments related by a quasi-two-fold rotational symmetry and a multidrug-binding cavity situated near the membrane surface. Here we report the crystal structure of an H(+)-coupled MATE transporter from Bacillus halodurans and the DinF subfamily at 3.2-Å resolution, unveiling a surprisingly asymmetric arrangement of 12 transmembrane helices. We also identified a membrane-embedded substrate-binding chamber by combining crystallographic and biochemical analyses. Our studies further suggested a direct competition between H(+) and substrate during DinF-mediated transport and implied how a MATE transporter alternates between its extracellular- and intracellular-facing conformations to propel multidrug extrusion. Collectively, our results demonstrated heretofore-unrecognized mechanistic diversity among MATE transporters.
- Department of Biochemistry and Molecular Biology, Rosalind Franklin University of Medicine and Science, North Chicago, Illinois, USA.
Organizational Affiliation: