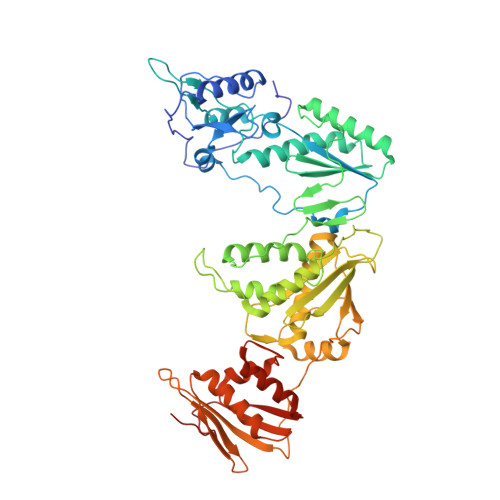
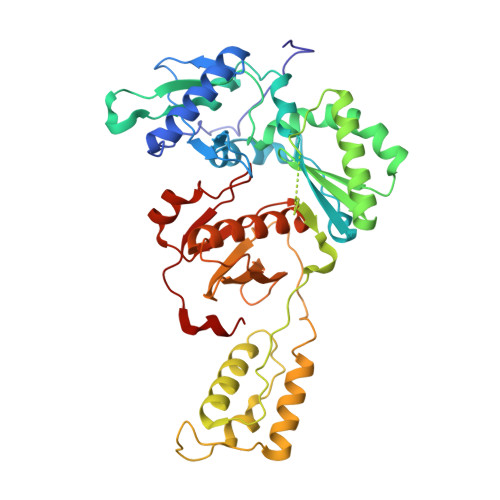
Structure-Based Evaluation of C5 Derivatives in the Catechol Diether Series Targeting HIV-1 Reverse Transcriptase.
Frey, K.M., Gray, W.T., Spasov, K.A., Bollini, M., Gallardo-Macias, R., Jorgensen, W.L., Anderson, K.S.(2014) Chem Biol Drug Des 83: 541-549
- PubMed: 24289305 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/cbdd.12266
- Primary Citation Related Structures:
4LSL, 4LSN - PubMed Abstract:
Using a computationally driven approach, a class of inhibitors with picomolar potency known as the catechol diethers were developed targeting the non-nucleoside-binding pocket of HIV-1 reverse transcriptase. Computational studies suggested that halogen-bonding interactions between the C5 substituent of the inhibitor and backbone carbonyl of conserved residue Pro95 might be important. While the recently reported crystal structures of the reverse transcriptase complexes confirmed the interactions with the non-nucleoside-binding pocket, they revealed the lack of a halogen-bonding interaction with Pro95. To understand the effects of substituents at the C5 position, we determined additional crystal structures with 5-Br and 5-H derivatives. Using comparative structural analysis, we identified several conformations of the ethoxy uracil dependent on the strength of a van der Waals interaction with the Cγ of Pro95 and the C5 substitution. The 5-Cl and 5-F derivatives position the ethoxy uracil to make more hydrogen bonds, whereas the larger 5-Br and smaller 5-H position the ethoxy uracil to make fewer hydrogen bonds. EC50 values correlate with the trends observed in the crystal structures. The influence of C5 substitutions on the ethoxy uracil conformation may have strategic value, as future derivatives can possibly be modulated to gain additional hydrogen-bonding interactions with resistant variants of reverse transcriptase.
- Department of Pharmacology, Yale University, 333 Cedar Street, SHM B350, New Haven, CT, 06520-8066, USA.
Organizational Affiliation: