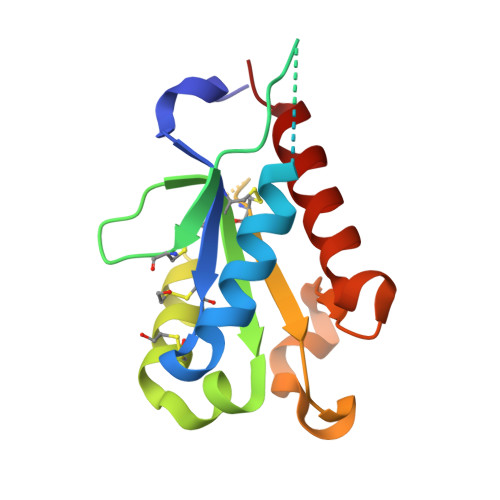
Crystal structures of the Toll/Interleukin-1 receptor (TIR) domains from the Brucella protein TcpB and host adaptor TIRAP reveal mechanisms of molecular mimicry.
Snyder, G.A., Deredge, D., Waldhuber, A., Fresquez, T., Wilkins, D.Z., Smith, P.T., Durr, S., Cirl, C., Jiang, J., Jennings, W., Luchetti, T., Snyder, N., Sundberg, E.J., Wintrode, P., Miethke, T., Xiao, T.S.(2014) J Biological Chem 289: 669-679
- PubMed: 24275656 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.523407
- Primary Citation Related Structures:
4LQC, 4LQD - PubMed Abstract:
The Toll/IL-1 receptor (TIR) domains are crucial innate immune signaling modules. Microbial TIR domain-containing proteins inhibit Toll-like receptor (TLR) signaling through molecular mimicry. The TIR domain-containing protein TcpB from Brucella inhibits TLR signaling through interaction with host adaptor proteins TIRAP/Mal and MyD88. To characterize the microbial mimicry of host proteins, we have determined the X-ray crystal structures of the TIR domains from the Brucella protein TcpB and the host adaptor protein TIRAP. We have further characterized homotypic interactions of TcpB using hydrogen/deuterium exchange mass spectrometry and heterotypic TcpB and TIRAP interaction by co-immunoprecipitation and NF-κB reporter assays. The crystal structure of the TcpB TIR domain reveals the microtubule-binding site encompassing the BB loop as well as a symmetrical dimer mediated by the DD and EE loops. This dimerization interface is validated by peptide mapping through hydrogen/deuterium exchange mass spectrometry. The human TIRAP TIR domain crystal structure reveals a unique N-terminal TIR domain fold containing a disulfide bond formed by Cys(89) and Cys(134). A comparison between the TcpB and TIRAP crystal structures reveals substantial conformational differences in the region that encompasses the BB loop. These findings underscore the similarities and differences in the molecular features found in the microbial and host TIR domains, which suggests mechanisms of bacterial mimicry of host signaling adaptor proteins, such as TIRAP.
- From the Laboratory of Immunology, NIAID, National Institutes of Health, Bethesda, Maryland 20892.
Organizational Affiliation: