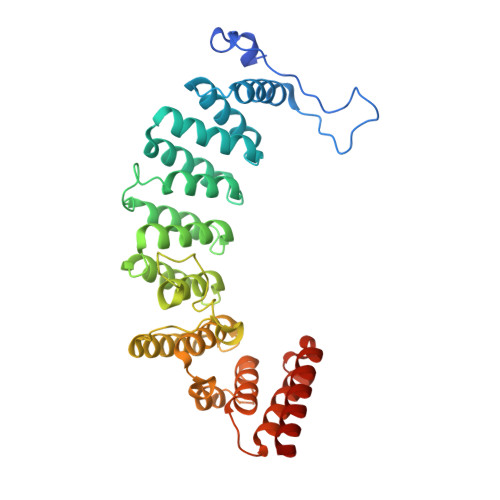
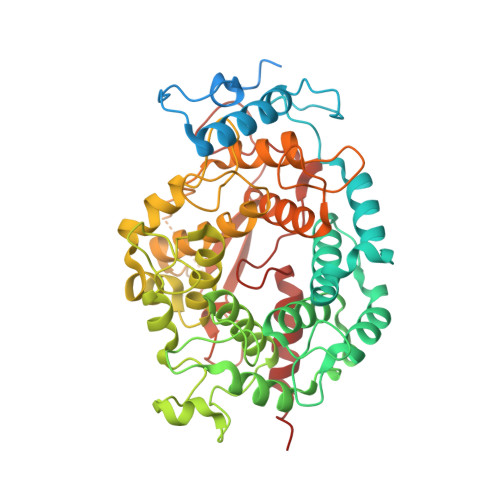
Crystal structures of the fungal pathogen Aspergillus fumigatus protein farnesyltransferase complexed with substrates and inhibitors reveal features for antifungal drug design.
Mabanglo, M.F., Hast, M.A., Lubock, N.B., Hellinga, H.W., Beese, L.S.(2014) Protein Sci 23: 289-301
- PubMed: 24347326 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2411
- Primary Citation Related Structures:
4L9P, 4LNB, 4LNG, 4MBG - PubMed Abstract:
Species of the fungal genus Aspergillus are significant human and agricultural pathogens that are often refractory to existing antifungal treatments. Protein farnesyltransferase (FTase), a critical enzyme in eukaryotes, is an attractive potential target for antifungal drug discovery. We report high-resolution structures of A. fumigatus FTase (AfFTase) in complex with substrates and inhibitors. Comparison of structures with farnesyldiphosphate (FPP) bound in the absence or presence of peptide substrate, corresponding to successive steps in ordered substrate binding, revealed that the second substrate-binding step is accompanied by motions of a loop in the catalytic site. Re-examination of other FTase structures showed that this motion is conserved. The substrate- and product-binding clefts in the AfFTase active site are wider than in human FTase (hFTase). Widening is a consequence of small shifts in the α-helices that comprise the majority of the FTase structure, which in turn arise from sequence variation in the hydrophobic core of the protein. These structural effects are key features that distinguish fungal FTases from hFTase. Their variation results in differences in steady-state enzyme kinetics and inhibitor interactions and presents opportunities for developing selective anti-fungal drugs by exploiting size differences in the active sites. We illustrate the latter by comparing the interaction of ED5 and Tipifarnib with hFTase and AfFTase. In AfFTase, the wider groove enables ED5 to bind in the presence of FPP, whereas in hFTase it binds only in the absence of substrate. Tipifarnib binds similarly to both enzymes but makes less extensive contacts in AfFTase with consequently weaker binding.
- Department of Biochemistry, Duke University Medical Center, Durham, North Carolina, 27710.
Organizational Affiliation: