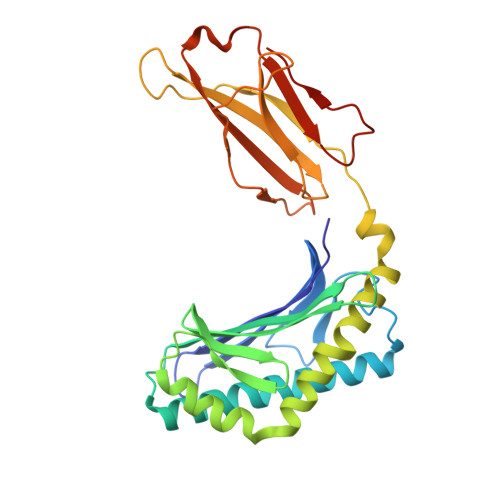
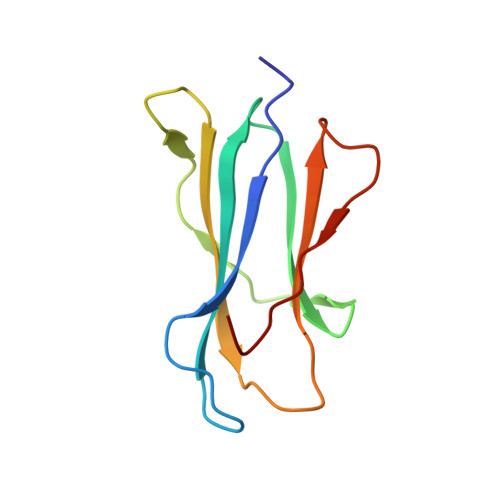
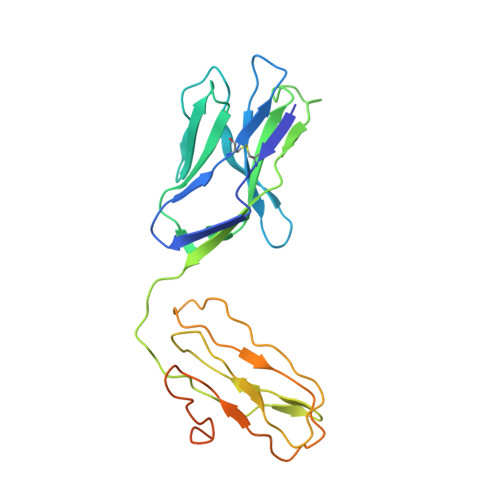
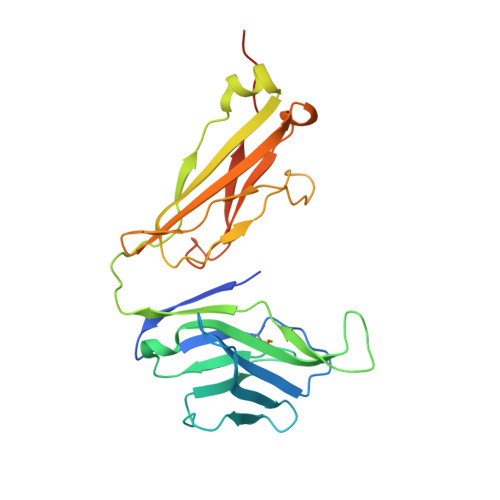
CD1d-lipid antigen recognition by the gamma delta TCR.
Uldrich, A.P., Le Nours, J., Pellicci, D.G., Gherardin, N.A., McPherson, K.G., Lim, R.T., Patel, O., Beddoe, T., Gras, S., Rossjohn, J., Godfrey, D.I.(2013) Nat Immunol 14: 1137-1145
- PubMed: 24076636 Search on PubMed
- DOI: https://doi.org/10.1038/ni.2713
- Primary Citation Related Structures:
4LFH, 4LHU - PubMed Abstract:
The T cell repertoire comprises αβ and γδ T cell lineages. Although it is established how αβ T cell antigen receptors (TCRs) interact with antigen presented by antigen-presenting molecules, this is unknown for γδ TCRs. We describe a population of human Vδ1(+) γδ T cells that exhibit autoreactivity to CD1d and provide a molecular basis for how a γδ TCR binds CD1d-α-galactosylceramide (α-GalCer). The γδ TCR docked orthogonally, over the A' pocket of CD1d, in which the Vδ1-chain, and in particular the germ line-encoded CDR1δ loop, dominated interactions with CD1d. The TCR γ-chain sat peripherally to the interface, with the CDR3γ loop representing the principal determinant for α-GalCer specificity. Accordingly, we provide insight into how a γδ TCR binds specifically to a lipid-loaded antigen-presenting molecule.
- 1] Department of Microbiology and Immunology, Peter Doherty Institute for Infection and Immunity, University of Melbourne, Parkville, Victoria, Australia. [2].
Organizational Affiliation: