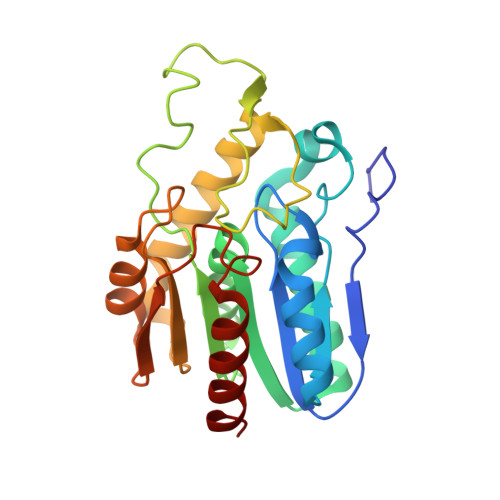
Substrate selectivity of bacterial monoacylglycerol lipase based on crystal structure
Tsurumura, T., Tsuge, H.(2014) J Struct Funct Genomics 15: 83-89
- PubMed: 24894647 Search on PubMed
- DOI: https://doi.org/10.1007/s10969-014-9181-2
- Primary Citation Related Structures:
4LHE - PubMed Abstract:
Lipases, which are conserved from bacteria to mammals, catalyze the hydrolysis of acylglycerol to free fatty acids and glycerol. Monoacylglycerol lipase (MGL) specifically catalyzes the hydrolysis of monoacylglycerol. Although there have been numerous studies of the structure of lipases, there have been few studies of MGL. Here, we report the crystal structure of authentic MGL isolated from Bacillus sp. H257 (bMGL). The crystal diffracts to 1.96 Å resolution. It belongs to space group P21212, and the unit cell parameters are a=99.7 Å, b=106.1 Å and c=43.0 Å. As in other lipases, three structural features for lipase activity are conserved in bMGL: the glycine-X-serine-X-glycine motif, catalytic triad and cap region. The structure of bMGL appears to be closed, as the cap region covers the active site entrance. The isolated bMGL hydrolyzed 2-AG, a known human MGL-specific substrate. Based on a 2-AG bound model, we discuss the substrate selectivity. The functional and structural features of bMGL provide insight how its substrate selectivity is determined and how specific inhibitors of bacterial MGL could be designed, which may be useful for development of novel antibiotics.
- Faculty of Life Sciences, Kyoto Sangyo University, Kamigamo-Motoyama, Kita-ku, Kyoto, 603-8555, Japan, ttsuru@cc.kyoto-su.ac.jp.
Organizational Affiliation: