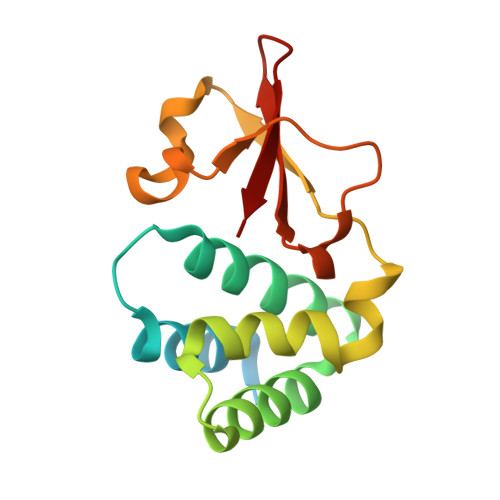
Ebolavirus VP35 coats the backbone of double-stranded RNA for interferon antagonism.
Bale, S., Julien, J.P., Bornholdt, Z.A., Krois, A.S., Wilson, I.A., Saphire, E.O.(2013) J Virol 87: 10385-10388
- PubMed: 23824825 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01452-13
- Primary Citation Related Structures:
4LG2 - PubMed Abstract:
Recognition of viral double-stranded RNA (dsRNA) activates interferon production and immune signaling in host cells. Crystal structures of ebolavirus VP35 show that it caps dsRNA ends to prevent sensing by pattern recognition receptors such as RIG-I. In contrast, structures of marburgvirus VP35 show that it primarily coats the dsRNA backbone. Here, we demonstrate that ebolavirus VP35 also coats the dsRNA backbone in solution, although binding to the dsRNA ends probably constitutes the initial binding event.
- Department of Immunology and Microbial Science, The Scripps Research Institute, La Jolla, California, USA.
Organizational Affiliation: