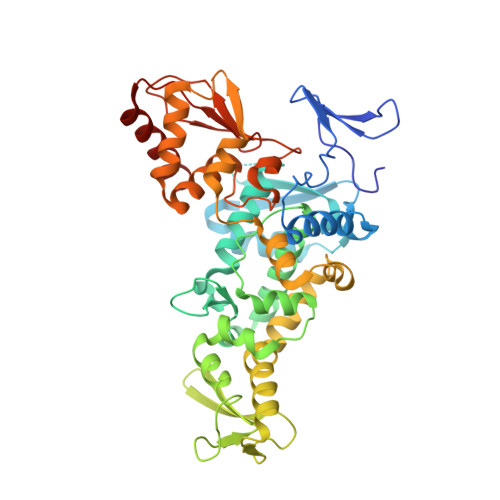
Mechanism of ubiquitin ligation and lysine prioritization by a HECT E3.
Kamadurai, H.B., Qiu, Y., Deng, A., Harrison, J.S., Macdonald, C., Actis, M., Rodrigues, P., Miller, D.J., Souphron, J., Lewis, S.M., Kurinov, I., Fujii, N., Hammel, M., Piper, R., Kuhlman, B., Schulman, B.A.(2013) Elife 2: e00828-e00828
- PubMed: 23936628 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.00828
- Primary Citation Related Structures:
4LCD - PubMed Abstract:
Ubiquitination by HECT E3 enzymes regulates myriad processes, including tumor suppression, transcription, protein trafficking, and degradation. HECT E3s use a two-step mechanism to ligate ubiquitin to target proteins. The first step is guided by interactions between the catalytic HECT domain and the E2∼ubiquitin intermediate, which promote formation of a transient, thioester-bonded HECT∼ubiquitin intermediate. Here we report that the second step of ligation is mediated by a distinct catalytic architecture established by both the HECT E3 and its covalently linked ubiquitin. The structure of a chemically trapped proxy for an E3∼ubiquitin-substrate intermediate reveals three-way interactions between ubiquitin and the bilobal HECT domain orienting the E3∼ubiquitin thioester bond for ligation, and restricting the location of the substrate-binding domain to prioritize target lysines for ubiquitination. The data allow visualization of an E2-to-E3-to-substrate ubiquitin transfer cascade, and show how HECT-specific ubiquitin interactions driving multiple reactions are repurposed by a major E3 conformational change to promote ligation. DOI:http://dx.doi.org/10.7554/eLife.00828.001.
- Department of Structural Biology , St Jude Children's Research Hospital , Memphis , United States.
Organizational Affiliation: