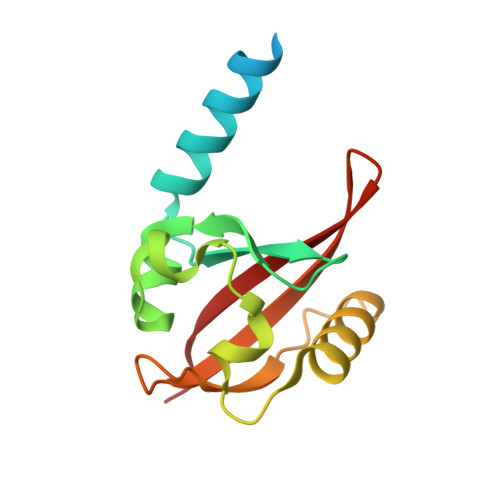
Multi-PAS domain-mediated protein oligomerization of PpsR from Rhodobacter sphaeroides.
Heintz, U., Meinhart, A., Winkler, A.(2014) Acta Crystallogr D Biol Crystallogr 70: 863-876
- PubMed: 24598755 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004713033634
- Primary Citation Related Structures:
4L9E, 4L9F, 4L9G - PubMed Abstract:
Per-ARNT-Sim (PAS) domains are essential modules of many multi-domain signalling proteins that mediate protein interaction and/or sense environmental stimuli. Frequently, multiple PAS domains are present within single polypeptide chains, where their interplay is required for protein function. Although many isolated PAS domain structures have been reported over the last decades, only a few structures of multi-PAS proteins are known. Therefore, the molecular mechanism of multi-PAS domain-mediated protein oligomerization and function is poorly understood. The transcription factor PpsR from Rhodobacter sphaeroides is such a multi-PAS domain protein that, in addition to its three PAS domains, contains a glutamine-rich linker and a C-terminal helix-turn-helix DNA-binding motif. Here, crystal structures of two N-terminally and C-terminally truncated PpsR variants that comprise a single (PpsRQ-PAS1) and two (PpsRN-Q-PAS1) PAS domains, respectively, are presented and the multi-step strategy required for the phasing of a triple PAS domain construct (PpsRΔHTH) is illustrated. While parts of the biologically relevant dimerization interface can already be observed in the two shorter constructs, the PpsRΔHTH structure reveals how three PAS domains enable the formation of multiple oligomeric states (dimer, tetramer and octamer), highlighting that not only the PAS cores but also their α-helical extensions are essential for protein oligomerization. The results demonstrate that the long helical glutamine-rich linker of PpsR results from a direct fusion of the N-cap of the PAS1 domain with the C-terminal extension of the N-domain that plays an important role in signal transduction.
- Department of Biomolecular Mechanisms, Max Planck Institute for Medical Research, Heidelberg, Germany.
Organizational Affiliation: