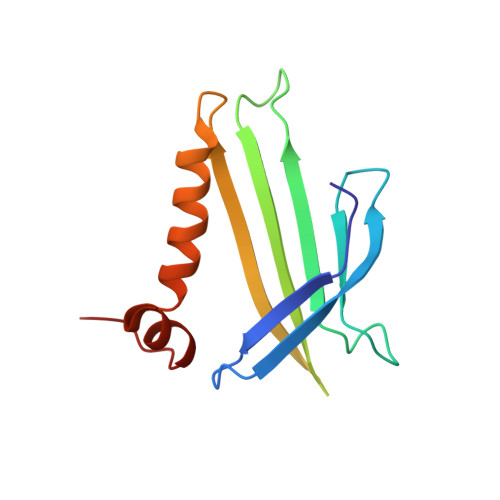
Crystal structure of the bacteriophage q beta coat protein in complex with the RNA operator of the replicase gene.
Rumnieks, J., Tars, K.(2014) J Mol Biology 426: 1039-1049
- PubMed: 24035813 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2013.08.025
- Primary Citation Related Structures:
4L8H - PubMed Abstract:
The coat proteins of single-stranded RNA bacteriophages specifically recognize and bind to a hairpin structure in their genome at the beginning of the replicase gene. The interaction serves to repress the synthesis of the replicase enzyme late in infection and contributes to the specific encapsidation of phage RNA. While this mechanism is conserved throughout the Leviviridae family, the coat protein and operator sequences from different phages show remarkable variation, serving as prime examples for the co-evolution of protein and RNA structure. To better understand the protein-RNA interactions in this virus family, we have determined the three-dimensional structure of the coat protein from bacteriophage Qβ bound to its cognate translational operator. The RNA binding mode of Qβ coat protein shares several features with that of the widely studied phage MS2, but only one nucleotide base in the hairpin loop makes sequence-specific contacts with the protein. Unlike in other RNA phages, the Qβ coat protein does not utilize an adenine-recognition pocket for binding a bulged adenine base in the hairpin stem but instead uses a stacking interaction with a tyrosine side chain to accommodate the base. The extended loop between β strands E and F of Qβ coat protein makes contacts with the lower part of the RNA stem, explaining the greater length dependence of the RNA helix for optimal binding to the protein. Consequently, the complex structure allows the proposal of a mechanism by which the Qβ coat protein recognizes and discriminates in favor of its cognate RNA.
- Biomedical Research and Study Center, Ratsupites 1, Riga LV1067, Latvia.
Organizational Affiliation: