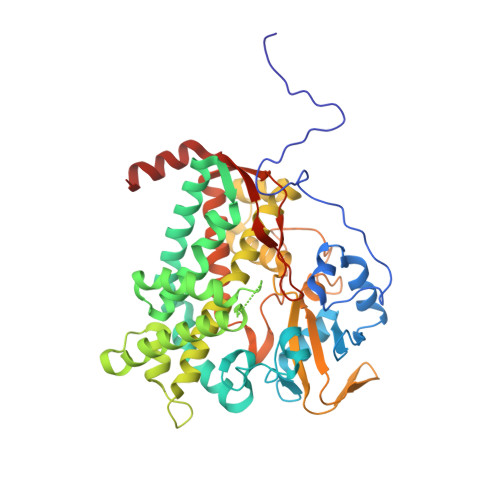
Cytochrome p450sky interacts directly with the nonribosomal Peptide synthetase to generate three amino Acid precursors in skyllamycin biosynthesis.
Uhlmann, S., Sussmuth, R.D., Cryle, M.J.(2013) ACS Chem Biol 8: 2586-2596
- PubMed: 24079328
- DOI: https://doi.org/10.1021/cb400555e
- Primary Citation Related Structures:
4L0E, 4L0F - PubMed Abstract:
The generation of modified amino acid precursors for incorporation in nonribosomal peptide synthesis (NRPS) plays a crucial, if often understated, role in the generation of peptide natural products. The biosynthesis of the cyclic depsipeptide skyllamycin requires three β-hydroxylated amino acid precursors, with in vivo gene inactivation experiments implicating cytochrome P450sky (CYP163B3) in the hydroxylation of these amino acids. Here, we demonstrate the in vitro oxidation of l-amino acid substrates bound to peptidyl carrier protein (PCP) domains 5, 7, and 11 of the skyllamycin nonribosomal synthetase by P450sky. Selectivity for these domains over other PCP domains could be demonstrated, with hydroxylation selective for l-amino acids and stereospecific in nature resulting in the (2S,3S)-configuration. The oxidation of amino acids or small molecule substrate analogues was not supported, demonstrating the necessity of the carrier protein in P450sky-catalyzed hydroxylation. The binding of aminoacyl-PCP substrates to P450sky was detected for the catalytically active PCP7 but not for the catalytically inactive PCP10, indicating carrier protein-mediated selectivity in P450sky substrate binding. X-ray crystal structures of P450sky reveal a 3D-structure with a highly open active site, the size of which is dictated by the carrier protein bound nature of the substrate. P450sky is the first P450 demonstrated to not only interact directly with PCP-bound amino acids within the peptide-forming NRPS but also to do so with three different PCP domains in a specific fashion. This represents an expansion of the complexity and scope of NRPS-mediated peptide synthesis, with the generation of hydroxylated amino acid precursors occurring through the interaction of P450 enzymes following, rather than prior to, the selection of amino acids by NRPS-adenylation domains.
- Institut für Chemie, Technische Universität Berlin , Strasse des 17. Juni 124, 10623 Berlin, Germany.
Organizational Affiliation: