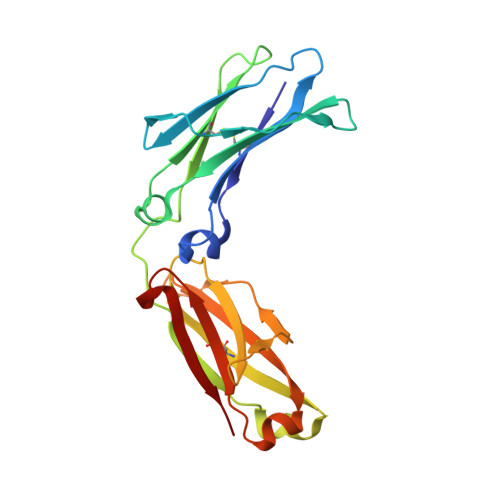
Immunoglobulin g1 fc domain motions: implications for fc engineering.
Frank, M., Walker, R.C., Lanzilotta, W.N., Prestegard, J.H., Barb, A.W.(2014) J Mol Biology 426: 1799-1811
- PubMed: 24522230 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2014.01.011
- Primary Citation Related Structures:
4KU1 - PubMed Abstract:
The fragment crystallizable (Fc) region links the key pathogen identification and destruction properties of immunoglobulin G (IgG). Pathogen opsonization positions Fcs to activate pro-inflammatory Fcγ receptors (FcγRs) on immune cells. The cellular response and committal to a damaging, though protective, immune response are tightly controlled at multiple levels. Control mechanisms are diverse and in many cases unclear, but one frequently suggested contribution originates in FcγR affinity being modulated through shifts in Fc conformational sampling. Here, we report a previously unseen IgG1 Fc conformation. This observation motivated an extensive molecular dynamics investigation of polypeptide and glycan motions that revealed greater amplitude of motion for the N-terminal Cγ2 domains and N-glycan than previously observed. Residues in the Cγ2/Cγ3 interface and disulfide-bonded hinge were identified as influencing the Cγ2 motion. Our results are consistent with a model of Fc that is structurally dynamic. Conformational states that are competent to bind immune-stimulating FcγRs interconverted with Fc conformations distinct from those observed in FcγR complexes, which may represent a transient, nonbinding population.
- Biognos AB, Generatorsgatan 1, 41705 Gothenburg, Sweden.
Organizational Affiliation: