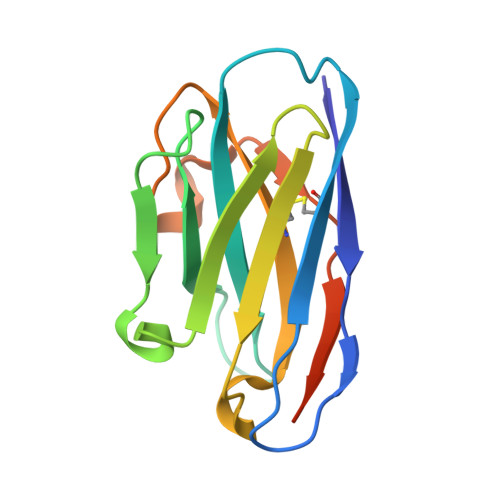
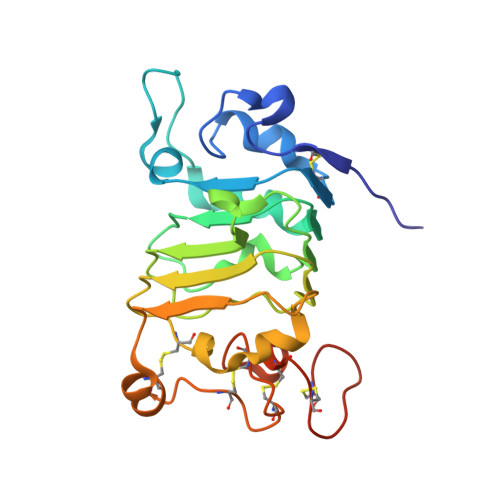
Structural Evaluation of EGFR Inhibition Mechanisms for Nanobodies/VHH Domains.
Schmitz, K.R., Bagchi, A., Roovers, R.C., van Bergen En Henegouwen, P.M.P., Ferguson, K.M.(2013) Structure 21: 1214-1224
- PubMed: 23791944 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2013.05.008
- Primary Citation Related Structures:
4KRL, 4KRM, 4KRN, 4KRO, 4KRP - PubMed Abstract:
The epidermal growth factor receptor (EGFR) is implicated in human cancers and is the target of several classes of therapeutic agents, including antibody-based drugs. Here, we describe X-ray crystal structures of the extracellular region of EGFR in complex with three inhibitory nanobodies, the variable domains of heavy chain only antibodies (VHH). VHH domains, the smallest natural antigen-binding modules, are readily engineered for diagnostic and therapeutic applications. All three VHH domains prevent ligand-induced EGFR activation, but use two distinct mechanisms. 7D12 sterically blocks ligand binding to EGFR in a manner similar to that of cetuximab. EgA1 and 9G8 bind an epitope near the EGFR domain II/III junction, preventing receptor conformational changes required for high-affinity ligand binding and dimerization. This epitope is accessible to the convex VHH paratope but inaccessible to the flatter paratope of monoclonal antibodies. Appreciating the modes of binding and inhibition of these VHH domains will aid in developing them for tumor imaging and/or cancer therapy.
- Department of Physiology and Graduate Group in Biochemistry and Molecular Biophysics, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA 19104, USA.
Organizational Affiliation: