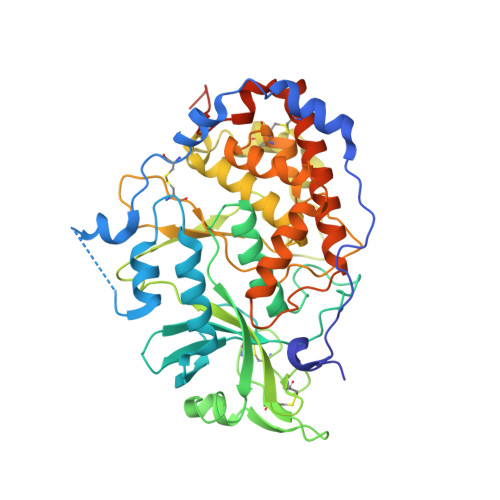
Crystal structure of the Golgi casein kinase.
Xiao, J., Tagliabracci, V.S., Wen, J., Kim, S.A., Dixon, J.E.(2013) Proc Natl Acad Sci U S A 110: 10574-10579
- PubMed: 23754375 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1309211110
- Primary Citation Related Structures:
4KQA, 4KQB - PubMed Abstract:
The family with sequence similarity 20 (Fam20) kinases phosphorylate extracellular substrates and play important roles in biomineralization. Fam20C is the Golgi casein kinase that phosphorylates secretory pathway proteins within Ser-x-Glu/pSer motifs. Mutations in Fam20C cause Raine syndrome, an osteosclerotic bone dysplasia. Here we report the crystal structure of the Fam20C ortholog from Caenorhabditis elegans. The nucleotide-free and Mn/ADP-bound structures unveil an atypical protein kinase-like fold and highlight residues critical for activity. The position of the regulatory αC helix and the lack of an activation loop indicate an architecture primed for efficient catalysis. Furthermore, several distinct elements, including the presence of disulfide bonds, suggest that the Fam20 family diverged early in the evolution of the protein kinase superfamily. Our results reinforce the structural diversity of protein kinases and have important implications for patients with disorders of biomineralization.
- Department of Pharmacology, University of California at San Diego, La Jolla, CA 92093, USA.
Organizational Affiliation: