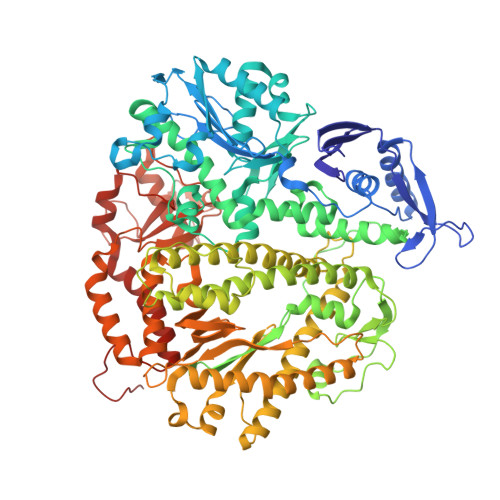
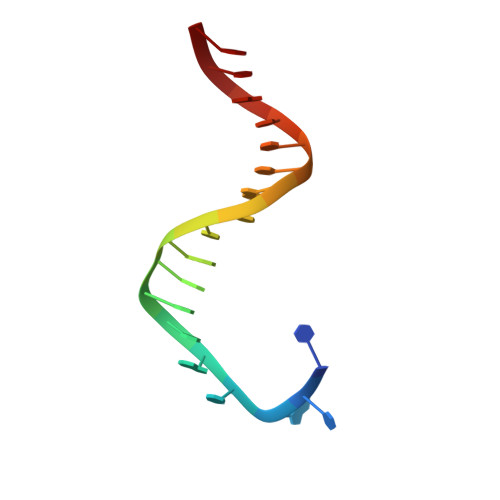
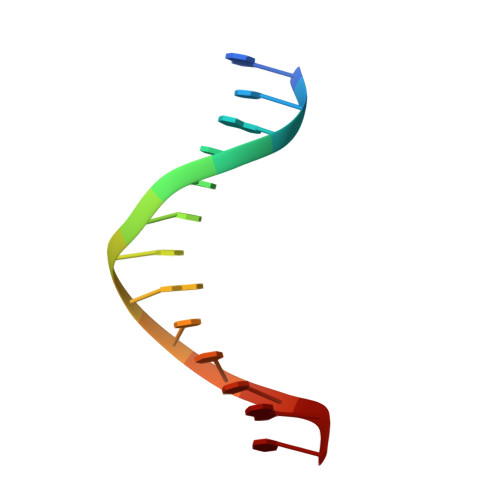
A Remote Palm Domain Residue of RB69 DNA Polymerase Is Critical for Enzyme Activity and Influences the Conformation of the Active Site.
Jacewicz, A., Trzemecka, A., Guja, K.E., Plochocka, D., Yakubovskaya, E., Bebenek, A., Garcia-Diaz, M.(2013) PLoS One 8: e76700-e76700
- PubMed: 24116139 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0076700
- Primary Citation Related Structures:
4I9L, 4I9Q, 4KHN - PubMed Abstract:
Non-conserved amino acids that are far removed from the active site can sometimes have an unexpected effect on enzyme catalysis. We have investigated the effects of alanine replacement of residues distant from the active site of the replicative RB69 DNA polymerase, and identified a substitution in a weakly conserved palm residue (D714A), that renders the enzyme incapable of sustaining phage replication in vivo. D714, located several angstroms away from the active site, does not contact the DNA or the incoming dNTP, and our apoenzyme and ternary crystal structures of the Pol(D714A) mutant demonstrate that D714A does not affect the overall structure of the protein. The structures reveal a conformational change of several amino acid side chains, which cascade out from the site of the substitution towards the catalytic center, substantially perturbing the geometry of the active site. Consistent with these structural observations, the mutant has a significantly reduced k pol for correct incorporation. We propose that the observed structural changes underlie the severe polymerization defect and thus D714 is a remote, non-catalytic residue that is nevertheless critical for maintaining an optimal active site conformation. This represents a striking example of an action-at-a-distance interaction.
- Department of Molecular Biology, Institute of Biochemistry and Biophysics, Polish Academy of Sciences, Warsaw, Poland.
Organizational Affiliation: