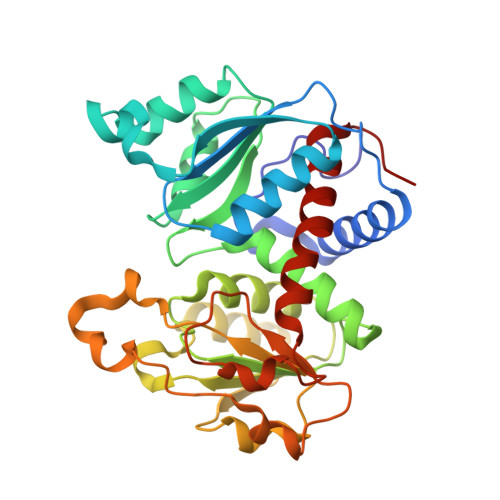
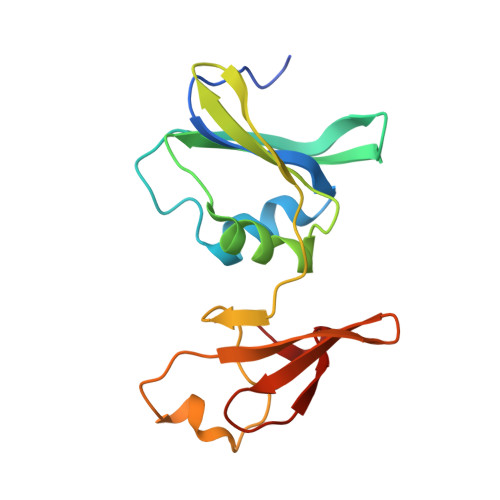
New Paradigm for Allosteric Regulation of Escherichia coli Aspartate Transcarbamoylase.
Cockrell, G.M., Zheng, Y., Guo, W., Peterson, A.W., Truong, J.K., Kantrowitz, E.R.(2013) Biochemistry 52: 8036-8047
- PubMed: 24138583 Search on PubMed
- DOI: https://doi.org/10.1021/bi401205n
- Primary Citation Related Structures:
4KGV, 4KGX, 4KGZ, 4KH0, 4KH1 - PubMed Abstract:
For nearly 60 years, the ATP activation and the CTP inhibition of Escherichia coli aspartate transcarbamoylase (ATCase) has been the textbook example of allosteric regulation. We present kinetic data and five X-ray structures determined in the absence and presence of a Mg(2+) concentration within the physiological range. In the presence of 2 mM divalent cations (Mg(2+), Ca(2+), Zn(2+)), CTP does not significantly inhibit the enzyme, while the allosteric activation by ATP is enhanced. The data suggest that the actual allosteric inhibitor of ATCase in vivo is the combination of CTP, UTP, and a divalent cation, and the actual allosteric activator is a divalent cation with ATP or ATP and GTP. The structural data reveals that two NTPs can bind to each allosteric site with a divalent cation acting as a bridge between the triphosphates. Thus, the regulation of ATCase is far more complex than previously believed and calls many previous studies into question. The X-ray structures reveal that the catalytic chains undergo essentially no alternations; however, several regions of the regulatory chains undergo significant structural changes. Most significant is that the N-terminal region of the regulatory chains exists in different conformations in the allosterically activated and inhibited forms of the enzyme. Here, a new model of allosteric regulation is proposed.
- Department of Chemistry, Boston College , Merkert Chemistry Center, 2609 Beacon Street, Chestnut Hill, MA 02467 U.S.A.
Organizational Affiliation: