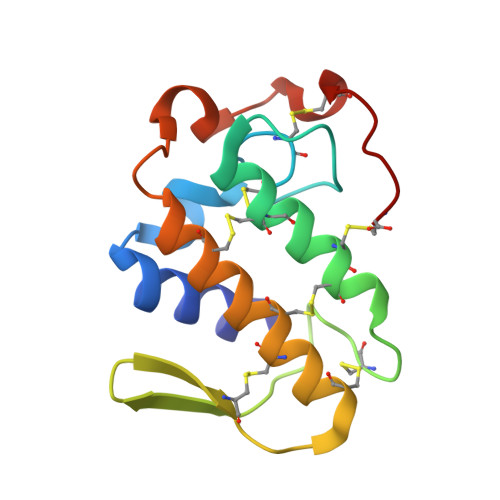
Structural and functional studies with mytoxin II from Bothrops moojeni reveal remarkable similarities and differences compared to other catalytically inactive phospholipases A2-like.
Salvador, G.H., Cavalcante, W.L., Dos Santos, J.I., Gallacci, M., Soares, A.M., Fontes, M.R.(2013) Toxicon 72C: 52-63
- PubMed: 23810946 Search on PubMed
- DOI: https://doi.org/10.1016/j.toxicon.2013.06.013
- Primary Citation Related Structures:
4KF3 - PubMed Abstract:
Lys49-phospholipases A₂ (Lys49-PLA₂s) are proteins found in bothropic snake venoms (Viperidae family) and belong to a class of proteins which presents a phospholipase A2 scaffold but are catalytically inactive. These proteins (also known as PLA₂s-like toxins) exert a pronounced local myotoxic effect and are not neutralized by antivenom, being their study relevant in terms of medical and scientific interest. Despite of the several studies reported in the literature for this class of proteins only a partial consensus has been achieved concerning their functional-structural relationships. In this work, we present a comprehensive structural and functional study with the MjTX-II, a dimeric Lys49-PLA₂ from Bothrops moojeni venom which includes: (i) high-resolution crystal structure; (ii) dynamic light scattering and bioinformatics studies in order to confirm its biological assembly; (iii) myographic and electrophysiological studies and, (iv) comparative studies with other Lys49-PLA₂s. These comparative analyses let us to get important insights into the role of Lys122 amino acid, previously indicated as responsible for Lys49-PLA₂s catalytic inactivity and added important elements to establish the correct biological assembly for this class of proteins. Furthermore, we show two unique sequential features of MjTX-II (an amino acid insertion and a mutation) in comparison to all bothropic Lys49-PLA₂s that lead to a distinct way of ligand binding at the toxin's hydrophobic channel and also, allowed the presence of an additional ligand molecule in this region. These facts suggest a possible particular mode of binding for long-chain ligands that interacts with MjTX-II hydrophobic channel, a feature that may directly affect the design of structure-based ligands for Lys49-PLA₂s.
- Depto. de Física e Biofísica, Instituto de Biociências, UNESP-Univ Estadual Paulista, Botucatu, SP, Brazil.
Organizational Affiliation: