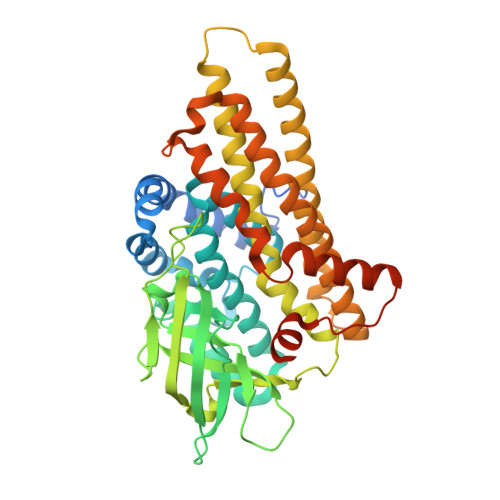
Active site architecture of a sugar N-oxygenase.
Thoden, J.B., Branch, M.C., Zimmer, A.L., Bruender, N.A., Holden, H.M.(2013) Biochemistry 52: 3191-3193
- PubMed: 23621882 Search on PubMed
- DOI: https://doi.org/10.1021/bi400407x
- Primary Citation Related Structures:
4KCF - PubMed Abstract:
KijD3 is a flavin-dependent N-oxygenase implicated in the formation of the nitro-containing sugar d-kijanose, found attached to the antibiotic kijanimicin. For this investigation, the structure of KijD3 in complex with FMN and its dTDP-sugar substrate was solved to 2.1 Å resolution. In contrast to the apoenzyme structure, the C-terminus of the protein becomes ordered and projects into the active site cleft [Bruender, N. A., Thoden, J. B., and Holden, H. M. (2010) Biochemistry 49, 3517-3524]. The amino group of the dTDP-aminosugar that is oxidized is located 4.9 Å from C4a of the flavin ring. The model provides a molecular basis for understanding the manner in which KijD3 catalyzes its unusual chemical transformation.
- Department of Biochemistry, University of Wisconsin , Madison, Wisconsin 53706, United States.
Organizational Affiliation: