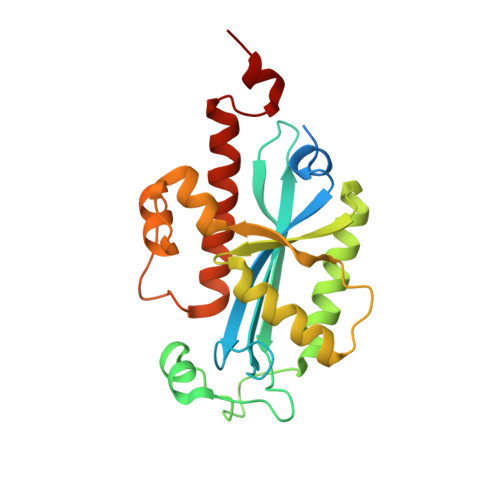
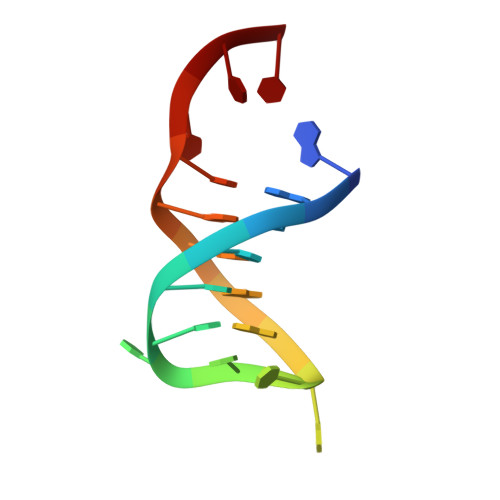
Structural insights into DNA repair by RNase T--an exonuclease processing 3' end of structured DNA in repair pathways.
Hsiao, Y.Y., Fang, W.H., Lee, C.C., Chen, Y.P., Yuan, H.S.(2014) PLoS Biol 12: e1001803-e1001803
- PubMed: 24594808 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.1001803
- Primary Citation Related Structures:
4KAZ, 4KB0, 4KB1 - PubMed Abstract:
DNA repair mechanisms are essential for preservation of genome integrity. However, it is not clear how DNA are selected and processed at broken ends by exonucleases during repair pathways. Here we show that the DnaQ-like exonuclease RNase T is critical for Escherichia coli resistance to various DNA-damaging agents and UV radiation. RNase T specifically trims the 3' end of structured DNA, including bulge, bubble, and Y-structured DNA, and it can work with Endonuclease V to restore the deaminated base in an inosine-containing heteroduplex DNA. Crystal structure analyses further reveal how RNase T recognizes the bulge DNA by inserting a phenylalanine into the bulge, and as a result the 3' end of blunt-end bulge DNA can be digested by RNase T. In contrast, the homodimeric RNase T interacts with the Y-structured DNA by a different binding mode via a single protomer so that the 3' overhang of the Y-structured DNA can be trimmed closely to the duplex region. Our data suggest that RNase T likely processes bulge and bubble DNA in the Endonuclease V-dependent DNA repair, whereas it processes Y-structured DNA in UV-induced and various other DNA repair pathways. This study thus provides mechanistic insights for RNase T and thousands of DnaQ-like exonucleases in DNA 3'-end processing.
- Institute of Molecular Biology, Academia Sinica, Taipei, Taiwan, Republic of China; Department of Biological Science and Technology, National Chiao Tung University, Hsinchu, Taiwan, Republic of China.
Organizational Affiliation: