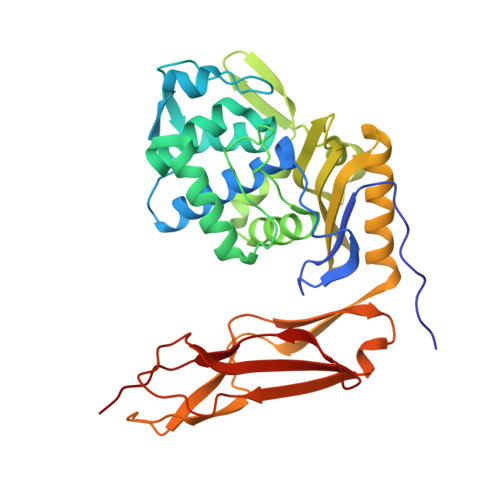
Structural analysis of the role of Pseudomonas aeruginosa penicillin-binding protein 5 in beta-lactam resistance.
Smith, J.D., Kumarasiri, M., Zhang, W., Hesek, D., Lee, M., Toth, M., Vakulenko, S., Fisher, J.F., Mobashery, S., Chen, Y.(2013) Antimicrob Agents Chemother 57: 3137-3146
- PubMed: 23629710 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AAC.00505-13
- Primary Citation Related Structures:
4K91 - PubMed Abstract:
Penicillin-binding protein 5 (PBP5) is one of the most abundant PBPs in Pseudomonas aeruginosa. Although its main function is that of a cell wall dd-carboxypeptidase, it possesses sufficient β-lactamase activity to contribute to the ability of P. aeruginosa to resist the antibiotic activity of the β-lactams. The study of these dual activities is important for understanding the mechanisms of antibiotic resistance by P. aeruginosa, an important human pathogen, and to the understanding of the evolution of β-lactamase activity from the PBP enzymes. We purified a soluble version of P. aeruginosa PBP5 (designated Pa sPBP5) by deletion of its C-terminal membrane anchor. Under in vitro conditions, Pa sPBP5 demonstrates both dd-carboxypeptidase and expanded-spectrum β-lactamase activities. Its crystal structure at a 2.05-Å resolution shows features closely resembling those of the class A β-lactamases, including a shortened loop spanning residues 74 to 78 near the active site and with respect to the conformations adopted by two active-site residues, Ser101 and Lys203. These features are absent in the related PBP5 of Escherichia coli. A comparison of the two Pa sPBP5 monomers in the asymmetric unit, together with molecular dynamics simulations, revealed an active-site flexibility that may explain its carbapenemase activity, a function that is absent in the E. coli PBP5 enzyme. Our functional and structural characterizations underscore the versatility of this PBP5 in contributing to the β-lactam resistance of P. aeruginosa while highlighting how broader β-lactamase activity may be encoded in the structural folds shared by the PBP and serine β-lactamase classes.
- Department of Molecular Medicine, University of South Florida, Tampa, Florida, USA.
Organizational Affiliation: