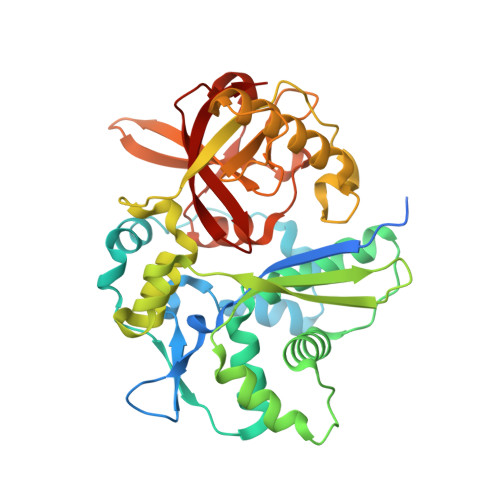
Structure and catalytic mechanism of yeast 4-amino-4-deoxychorismate lyase
Dai, Y.-N., Chi, C.-B., Zhou, K., Cheng, W., Jiang, Y.-L., Ren, Y.-M., Ruan, K., Chen, Y., Zhou, C.-Z.(2013) J Biological Chem 288: 22985-22992
- PubMed: 23818518
- DOI: https://doi.org/10.1074/jbc.M113.480335
- Primary Citation Related Structures:
4K6N - PubMed Abstract:
Saccharomyces cerevisiae Abz2 is a pyridoxal 5'-phosphate (PLP)-dependent lyase that converts 4-amino-4-deoxychorismate (ADC) to para-aminobenzoate and pyruvate. To investigate the catalytic mechanism, we determined the 1.9 Å resolution crystal structure of Abz2 complexed with PLP, representing the first eukaryotic ADC lyase structure. Unlike Escherichia coli ADC lyase, whose dimerization is critical to the formation of the active site, the overall structure of Abz2 displays as a monomer of two domains. At the interdomain cleft, a molecule of cofactor PLP forms a Schiff base with residue Lys-251. Computational simulations defined a basic clamp to orientate the substrate ADC in a proper pose, which was validated by site-directed mutageneses combined with enzymatic activity assays. Altogether, we propose a putative catalytic mechanism of a unique class of monomeric ADC lyases led by yeast Abz2.
- Hefei National Laboratory for Physical Sciences at the Microscale and School of Life Sciences, University of Science and Technology of China, Hefei Anhui 230027, China.
Organizational Affiliation: