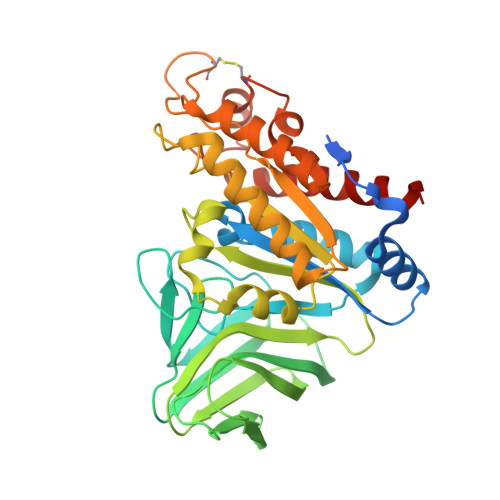
Visualization of a substrate-induced productive conformation of the catalytic triad of the Neisseria meningitidis peptidoglycan O-acetylesterase reveals mechanistic conservation in SGNH esterase family members.
Williams, A.H., Veyrier, F.J., Bonis, M., Michaud, Y., Raynal, B., Taha, M.K., White, S.W., Haouz, A., Boneca, I.G.(2014) Acta Crystallogr D Biol Crystallogr 70: 2631-2639
- PubMed: 25286847 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1399004714016770
- Primary Citation Related Structures:
4K3U, 4K40, 4K7J, 4K9S - PubMed Abstract:
Peptidoglycan O-acetylesterase (Ape1), which is required for host survival in Neisseria sp., belongs to the diverse SGNH hydrolase superfamily, which includes important viral and bacterial virulence factors. Here, multi-domain crystal structures of Ape1 with an SGNH catalytic domain and a newly identified putative peptidoglycan-detection module are reported. Enzyme catalysis was performed in Ape1 crystals and key catalytic intermediates along the SGNH esterase hydrolysis reaction pathway were visualized, revealing a substrate-induced productive conformation of the catalytic triad, a mechanistic detail that has not previously been observed. This substrate-induced productive conformation of the catalytic triad shifts the established dogma on these enzymes, generating valuable insight into the structure-based design of drugs targeting the SGNH esterase superfamily.
- Institut Pasteur, Unité Biologie et génétique de la paroi bactérienne, Dept. Microbiologie, 28 Rue du Dr. Roux, 75015 Paris, France.
Organizational Affiliation: