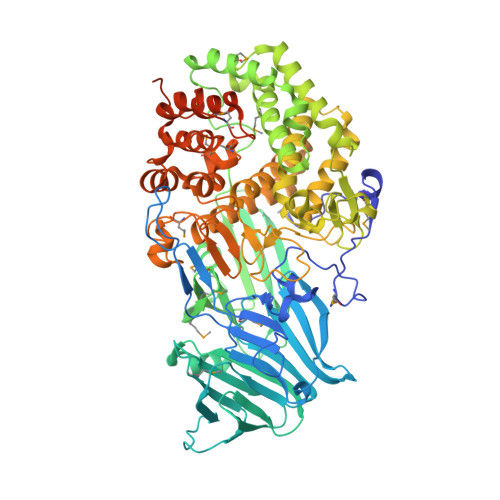
The structure of a glycoside hydrolase family 81 endo-[beta]-1,3-glucanase
Zhou, P., Chen, Z.Z., Yan, Q.J., Yang, S.Q., Hilgenfeld, R., Jiang, Z.Q.(2013) Acta Crystallogr D Biol Crystallogr 69: 2027-2038
- PubMed: 18802694 Search on PubMed
- DOI: https://doi.org/10.1007/s00253-008-1617-9
- Primary Citation Related Structures:
4K35, 4K3A - PubMed Abstract:
A beta-1,3-glucanase gene, encoding a protein of 1,793 amino acids, was cloned from a strain of Paenibacillus sp. in this study. This large protein, designated as LamA, consists of many putative functional units, which include, from N to C terminus, a leader peptide, three repeats of the S-layer homologous module, a catalytic module of glycoside hydrolase family 16, four repeats of the carbohydrate-binding module of family CBM_4_9, and an analogue of coagulation factor Fa5/8C. Several truncated proteins, composed of the catalytic module with various organizations of the appended modules, were successfully expressed and characterized in this study. Data indicated that the catalytic module specifically hydrolyze beta-1,3- and beta-1,3-1,4-glucans. Also, laminaritriose was the major product upon endolytic hydrolysis of laminarin. The CBM repeats and Fa5/8C analogue substantially enhanced the hydrolyzing activity of the catalytic module, particularly toward insoluble complex substrates, suggesting their modulating functions in the enzymatic activity of LamA. Carbohydrate-binding assay confirmed the binding capabilities of the CBM repeats and Fa5/8C analogue to beta-1,3-, beta-1,3-1,4-, and even beta-1,4-glucans. These appended modules also enhanced the inhibition effect of the catalytic module on the growth of Candida albicans and Rhizoctonia solani.
- Graduate Institute of Biotechnology, National Chung Hsing University, 250 Kuo-Kuang Rd, Taichung, Taiwan 40227, Republic of China.
Organizational Affiliation: