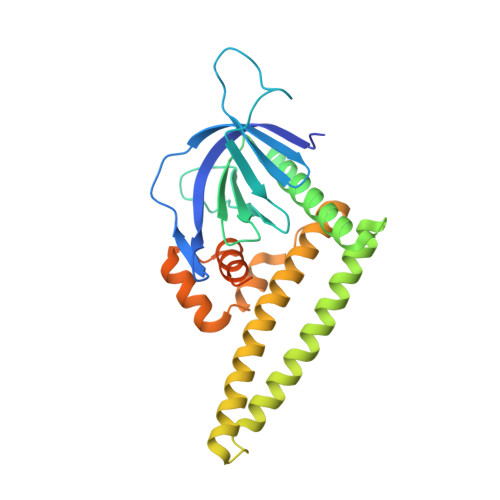
High-resolution structure of the Tiam1 PHn-CC-Ex domain.
Joshi, M., Gakhar, L., Fuentes, E.J.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 744-752
- PubMed: 23832200 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113014206
- Primary Citation Related Structures:
4K2O, 4K2P - PubMed Abstract:
The T-lymphoma and metastasis gene 1 (TIAM1) encodes a guanine nucleotide-exchange factor protein (Tiam1) that is specific for the Rho-family GTPase Rac1 and is important for cell polarity, migration and adhesion. Tiam1 is a large multi-domain protein that contains several protein-protein binding domains that are important for regulating cellular function. The PHn-CC-Ex domain is critical for plasma-membrane association and interactions with protein-scaffold proteins (e.g. Par3b, spinophilin, IRSp53 and JIP2) that direct Tiam1-Rac1 signaling specificity. It was determined that the coiled-coil domain of Par3b binds the PHn-CC-Ex domain with a dissociation constant of ≈ 30 µM. Moreover, the structures of two variants of the Tiam1 PHn-CC-Ex domain were solved at resolutions of 1.98 and 2.15 Å, respectively. The structures indicate that the PHn, CC and Ex regions form independent subdomains that together provide an integrated platform for binding partner proteins. Small-angle X-ray scattering (SAXS) data indicate that the Tiam1 PHn-CC-Ex domain is monomeric in solution and that the solution and crystal structures are very similar. Together, these data provide the foundation necessary to elucidate the structural mechanism of the PHn-CC-Ex/scaffold interactions that are critical for Tiam1-Rac1 signaling specificity.
- Department of Biochemistry, Roy J. and Lucille A. Carver College of Medicine, University of Iowa, Iowa City, IA 52242-1109, USA.
Organizational Affiliation: