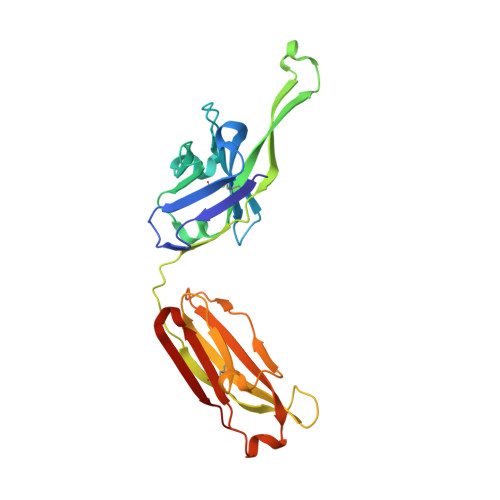
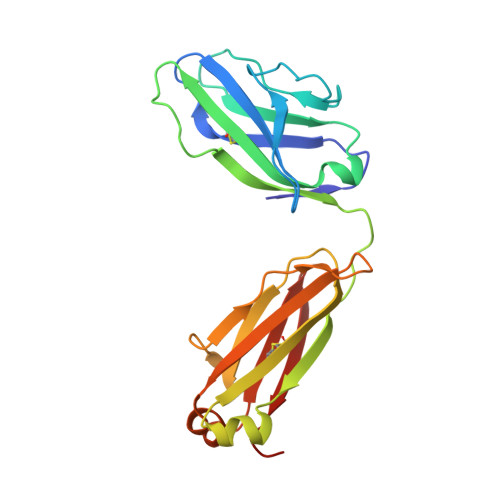
Broadly neutralizing antibody PGT121 allosterically modulates CD4 binding via recognition of the HIV-1 gp120 V3 base and multiple surrounding glycans.
Julien, J.P., Sok, D., Khayat, R., Lee, J.H., Doores, K.J., Walker, L.M., Ramos, A., Diwanji, D.C., Pejchal, R., Cupo, A., Katpally, U., Depetris, R.S., Stanfield, R.L., McBride, R., Marozsan, A.J., Paulson, J.C., Sanders, R.W., Moore, J.P., Burton, D.R., Poignard, P., Ward, A.B., Wilson, I.A.(2013) PLoS Pathog 9: e1003342-e1003342
- PubMed: 23658524 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1003342
- Primary Citation Related Structures:
4JY4, 4JY5, 4JY6 - PubMed Abstract:
New broad and potent neutralizing HIV-1 antibodies have recently been described that are largely dependent on the gp120 N332 glycan for Env recognition. Members of the PGT121 family of antibodies, isolated from an African donor, neutralize ∼70% of circulating isolates with a median IC50 less than 0.05 µg ml(-1). Here, we show that three family members, PGT121, PGT122 and PGT123, have very similar crystal structures. A long 24-residue HCDR3 divides the antibody binding site into two functional surfaces, consisting of an open face, formed by the heavy chain CDRs, and an elongated face, formed by LCDR1, LCDR3 and the tip of the HCDR3. Alanine scanning mutagenesis of the antibody paratope reveals a crucial role in neutralization for residues on the elongated face, whereas the open face, which accommodates a complex biantennary glycan in the PGT121 structure, appears to play a more secondary role. Negative-stain EM reconstructions of an engineered recombinant Env gp140 trimer (SOSIP.664) reveal that PGT122 interacts with the gp120 outer domain at a more vertical angle with respect to the top surface of the spike than the previously characterized antibody PGT128, which is also dependent on the N332 glycan. We then used ITC and FACS to demonstrate that the PGT121 antibodies inhibit CD4 binding to gp120 despite the epitope being distal from the CD4 binding site. Together, these structural, functional and biophysical results suggest that the PGT121 antibodies may interfere with Env receptor engagement by an allosteric mechanism in which key structural elements, such as the V3 base, the N332 oligomannose glycan and surrounding glycans, including a putative V1/V2 complex biantennary glycan, are conformationally constrained.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, California, USA.
Organizational Affiliation: