Ring-Opening of the gamma-OH-PdG Adduct Promotes Error-Free Bypass by the Sulfolobus solfataricus DNA Polymerase Dpo4.
Shanmugam, G., Minko, I.G., Banerjee, S., Christov, P.P., Kozekov, I.D., Rizzo, C.J., Lloyd, R.S., Egli, M., Stone, M.P.(2013) Chem Res Toxicol 26: 1348-1360
- PubMed: 23947567 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/tx400200b
- Primary Citation Related Structures:
4JUZ, 4JV0, 4JV1, 4JV2 - PubMed Abstract:
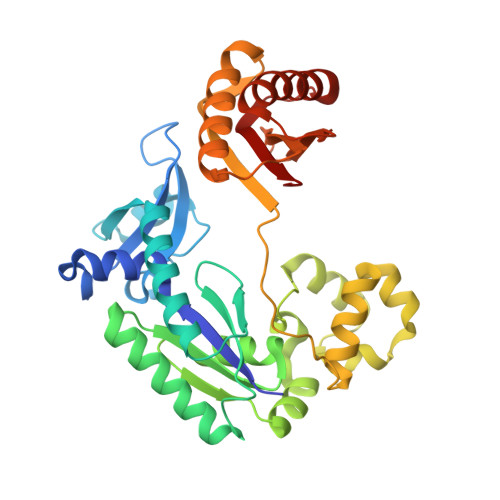
Acrolein, a mutagenic aldehyde, reacts with deoxyguanosine (dG) to form 3-(2'-deoxy-β-d-erythro-pentofuranosyl)-5,6,7,8-tetrahydro-8-hydroxypyrimido[1,2-a] purin-10(3H)-one (γ-OH-PdG). When placed opposite deoxycytosine (dC) in DNA, γ-OH-PdG undergoes ring-opening to the N(2)-(3-oxopropyl)-dG. Ring-opening of the adduct has been hypothesized to facilitate nonmutagenic bypass, particularly by DNA polymerases of the Y family. This study examined the bypass of γ-OH-PdG by Sulfolobus solfataricus Dpo4, the prototypic Y-family DNA polymerase, using templates that contained the adduct in either the 5'-CXG-3' or the 5'-TXG-3' sequence context. Although γ-OH-PdG partially blocked Dpo4-catalyzed DNA synthesis, full primer extension was observed, and the majority of bypass products were error-free. Conversion of the adduct into an irreversibly ring-opened derivative prior to reaction facilitated bypass and further improved the fidelity. Structures of ternary Dpo4·DNA·dNTP complexes were determined with primers that either were positioned immediately upstream of the lesion (preinsertion complexes) or had a 3'-terminal dC opposite the lesion (postinsertion complexes); the incoming nucleotides, either dGTP or dATP, were complementary to the template 5'-neighbor nucleotide. In both postinsertion complexes, the adduct existed as ring-opened species, and the resulting base-pair featured Watson-Crick hydrogen bonding. The incoming nucleotide paired with the 5'-neighbor template, while the primer 3'-hydroxyl was positioned to facilitate extension. In contrast, γ-OH-PdG was in the ring-closed form in both preinsertion complexes, and the overall structure did not favor catalysis. These data provide insights into γ-OH-PdG chemistry during replication bypass by the Dpo4 DNA polymerase and may explain why γ-OH-PdG-induced mutations due to primer-template misalignment are uncommon.
- Department of Chemistry, Center in Molecular Toxicology, Vanderbilt-Ingram Cancer Center, Vanderbilt Institute of Chemical Biology, and Center for Structural Biology, Vanderbilt University , Nashville, Tennessee 37235, United States.
Organizational Affiliation: