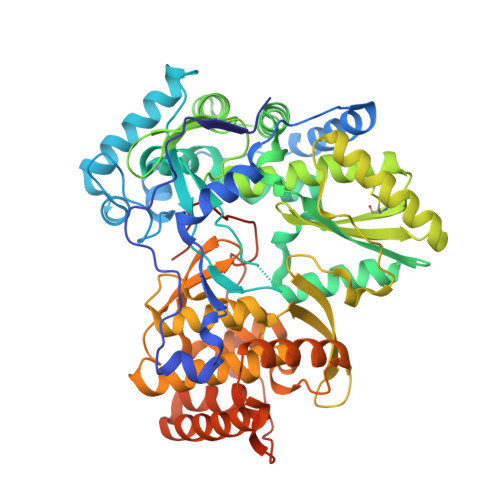
Molecular Dynamics Simulations and Structure-Based Rational Design Lead to Allosteric HCV NS5B Polymerase Thumb Pocket 2 Inhibitor with Picomolar Cellular Replicon Potency.
Hucke, O., Coulombe, R., Bonneau, P., Bertrand-Laperle, M., Brochu, C., Gillard, J., Joly, M.A., Landry, S., Lepage, O., Llinas-Brunet, M., Pesant, M., Poirier, M., Poirier, M., McKercher, G., Marquis, M., Kukolj, G., Beaulieu, P.L., Stammers, T.A.(2014) J Med Chem 57: 1932-1943
- PubMed: 23773186
- DOI: https://doi.org/10.1021/jm4004522
- Primary Citation Related Structures:
4JTW, 4JTY, 4JTZ, 4JU1, 4JU2 - PubMed Abstract:
The design and preliminary SAR of a new series of 1H-quinazolin-4-one (QAZ) allosteric HCV NS5B thumb pocket 2 (TP-2) inhibitors was recently reported. To support optimization efforts, a molecular dynamics (MD) based modeling workflow was implemented, providing information on QAZ binding interactions with NS5B. This approach predicted a small but critical ligand-binding induced movement of a protein backbone region which increases the pocket size and improves access to the backbone carbonyl groups of Val 494 and Pro 495. This localized backbone shift was consistent with key SAR results and was subsequently confirmed by X-ray crystallography. The MD protocol guided the design of inhibitors, exploiting novel H-bond interactions with the two backbone carbonyl groups, leading to the first thumb pocket 2 NS5B inhibitor with picomolar antiviral potency in genotype (gt) 1a and 1b replicons (EC50 = 120 and 110 pM, respectively) and with EC50 ≤ 80 nM against gt 2-6.
- Research and Development, Boehringer Ingelheim (Canada) Ltd. , 2100 Cunard Street, Laval, Quebec H7S 2G5, Canada.
Organizational Affiliation: