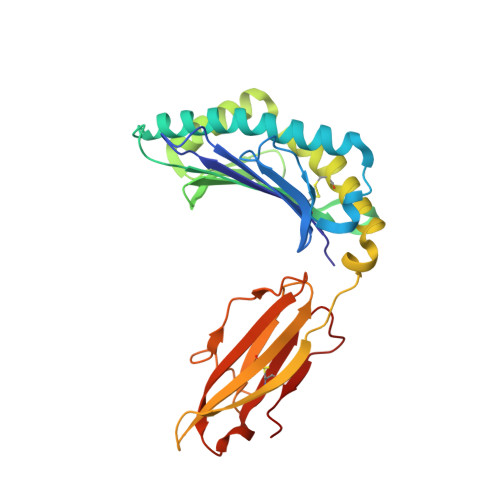
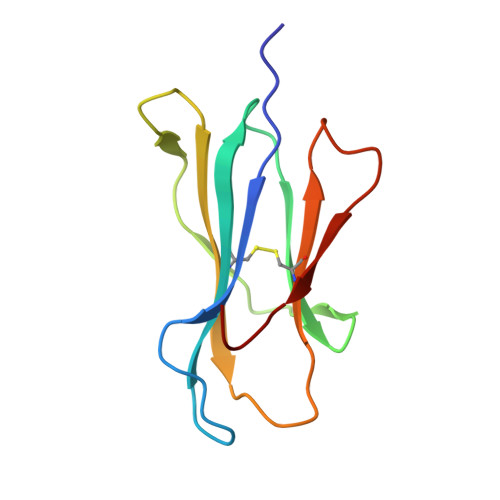
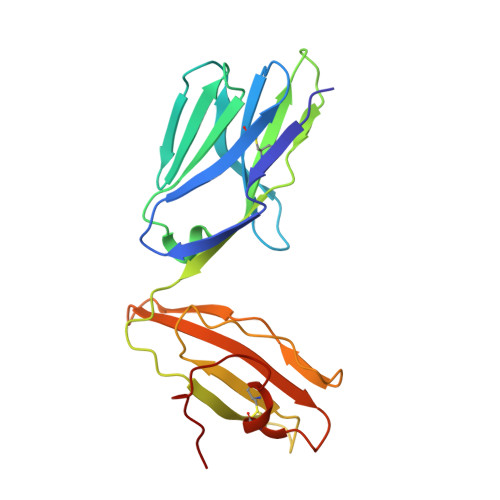
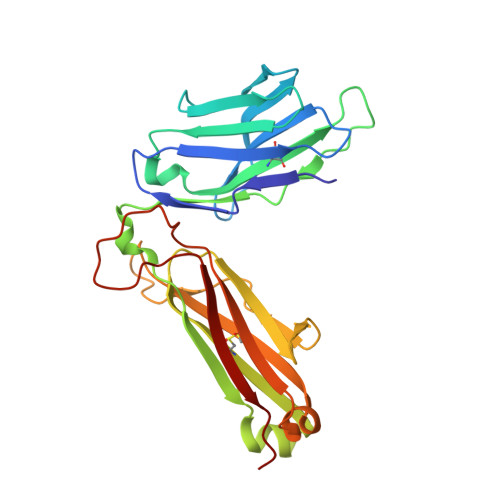
Highly divergent T-cell receptor binding modes underlie specific recognition of a bulged viral peptide bound to a human leukocyte antigen class I molecule.
Liu, Y.C., Miles, J.J., Neller, M.A., Gostick, E., Price, D.A., Purcell, A.W., McCluskey, J., Burrows, S.R., Rossjohn, J., Gras, S.(2013) J Biological Chem 288: 15442-15454
- PubMed: 23569211 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M112.447185
- Primary Citation Related Structures:
4JRX, 4JRY - PubMed Abstract:
Human leukocyte antigen (HLA)-I molecules can present long peptides, yet the mechanisms by which T-cell receptors (TCRs) recognize featured pHLA-I landscapes are unclear. We compared the binding modes of three distinct human TCRs, CA5, SB27, and SB47, complexed with a "super-bulged" viral peptide (LPEPLPQGQLTAY) restricted by HLA-B*35:08. The CA5 and SB27 TCRs engaged HLA-B*35:08(LPEP) similarly, straddling the central region of the peptide but making limited contacts with HLA-B*35:08. Remarkably, the CA5 TCR did not contact the α1-helix of HLA-B*35:08. Differences in the CDR3β loop between the CA5 and SB27 TCRs caused altered fine specificities. Surprisingly, the SB47 TCR engaged HLA-B*35:08(LPEP) using a completely distinct binding mechanism, namely "bypassing" the bulged peptide and making extensive contacts with the extreme N-terminal end of HLA-B*35:08. This docking footprint included HLA-I residues not observed previously as TCR contact sites. The three TCRs exhibited differing patterns of alloreactivity toward closely related or distinct HLA-I allotypes. Thus, the human T-cell repertoire comprises a range of TCRs that can interact with "bulged" pHLA-I epitopes using unpredictable strategies, including the adoption of atypical footprints on the MHC-I.
- Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton 3800, Australia.
Organizational Affiliation: