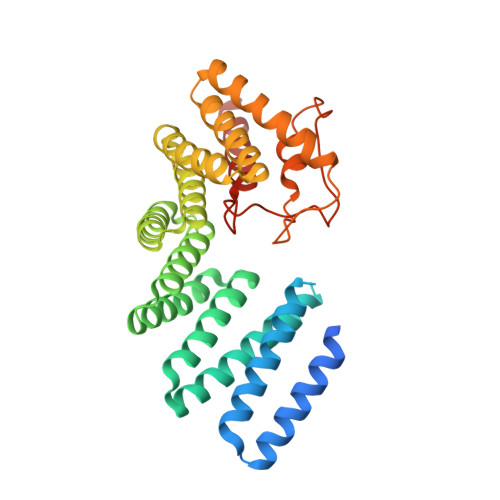
An autoinhibited conformation of LGN reveals a distinct interaction mode between GoLoco motifs and TPR motifs
Pan, Z., Zhu, J., Shang, Y., Wei, Z., Jia, M., Xia, C., Wen, W., Wang, W., Zhang, M.(2013) Structure 21: 1007-1017
- PubMed: 23665171 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2013.04.005
- Primary Citation Related Structures:
4JHR - PubMed Abstract:
LGN plays essential roles in asymmetric cell divisions via its N-terminal TPR-motif-mediated binding to mInsc and NuMA. This scaffolding activity requires the release of the autoinhibited conformation of LGN by binding of Gα(i) to its C-terminal GoLoco (GL) motifs. The interaction between the GL and TPR motifs of LGN represents a distinct GL/target binding mode with an unknown mechanism. Here, we show that two consecutive GL motifs of LGN form a minimal TPR-motif-binding unit. GL12 and GL34 bind to TPR0-3 and TPR4-7, respectively. The crystal structure of a truncated LGN reveals that GL34 forms a pair of parallel α helices and binds to the concave surface of TPR4-7, thereby preventing LGN from binding to other targets. Importantly, the GLs bind to TPR motifs with a mode distinct from that observed in the GL/Gα(i)·GDP complexes. Our results also indicate that multiple and orphan GL motif proteins likely respond to G proteins with distinct mechanisms.
- Shanghai Key Laboratory of Molecular Catalysis and Innovative Materials, Department of Chemistry and Institutes of Biomedical Sciences, Fudan University, Shanghai, China.
Organizational Affiliation: