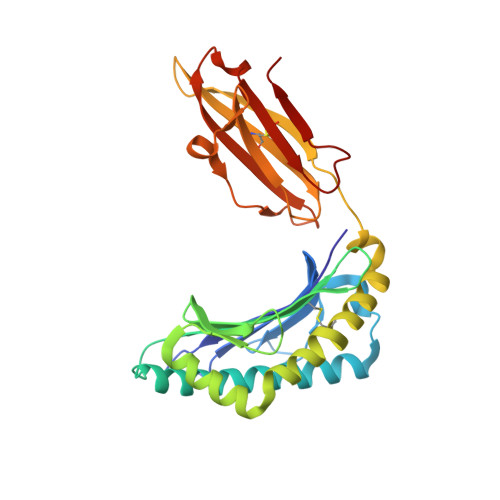
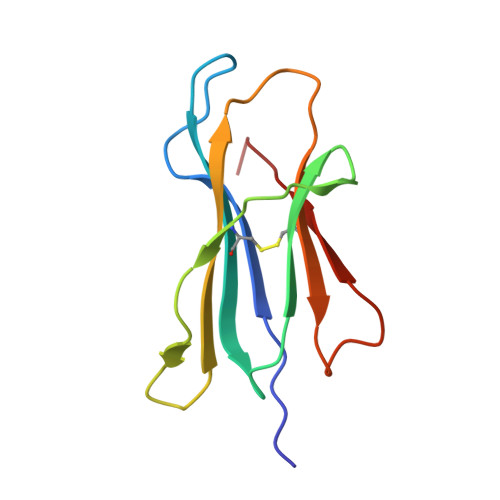
T-cell receptor specificity maintained by altered thermodynamics.
Madura, F., Rizkallah, P.J., Miles, K.M., Holland, C.J., Bulek, A.M., Fuller, A., Schauenburg, A.J., Miles, J.J., Liddy, N., Sami, M., Li, Y., Hossain, M., Baker, B.M., Jakobsen, B.K., Sewell, A.K., Cole, D.K.(2013) J Biological Chem 288: 18766-18775
- PubMed: 23698002 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M113.464560
- Primary Citation Related Structures:
4JFD, 4JFE, 4JFF, 4JFH, 4JFO, 4JFP, 4JFQ - PubMed Abstract:
The T-cell receptor (TCR) recognizes peptides bound to major histocompatibility molecules (MHC) and allows T-cells to interrogate the cellular proteome for internal anomalies from the cell surface. The TCR contacts both MHC and peptide in an interaction characterized by weak affinity (KD = 100 nM to 270 μM). We used phage-display to produce a melanoma-specific TCR (α24β17) with a 30,000-fold enhanced binding affinity (KD = 0.6 nM) to aid our exploration of the molecular mechanisms utilized to maintain peptide specificity. Remarkably, although the enhanced affinity was mediated primarily through new TCR-MHC contacts, α24β17 remained acutely sensitive to modifications at every position along the peptide backbone, mimicking the specificity of the wild type TCR. Thermodynamic analyses revealed an important role for solvation in directing peptide specificity. These findings advance our understanding of the molecular mechanisms that can govern the exquisite peptide specificity characteristic of TCR recognition.
- Cardiff University School of Medicine, Heath Park, Cardiff CF14 4XN, United Kingdom.
Organizational Affiliation: