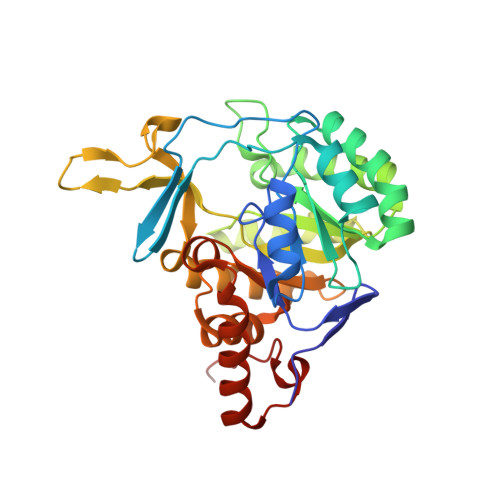
Structure of Trypanosoma cruzi dihydroorotate dehydrogenase in complex with MII-5-005
Inaoka, D.K., Iida, M., Tabuchi, T., Lee, N., Hashimoto, S., Matsuoka, S., Kuranaga, T., Shiba, T., Sakamoto, K., Suzuki, S., Balogun, E.O., Nara, T., Aoki, T., Inoue, M., Honma, T., Tanaka, A., Harada, S., Kita, K.To be published.