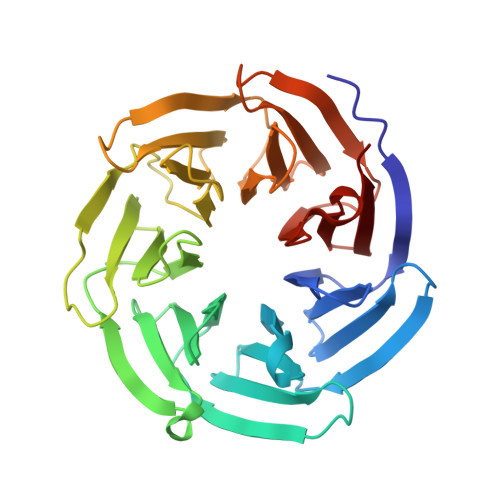
Rules for the recognition of dilysine retrieval motifs by coatomer.
Ma, W., Goldberg, J.(2013) EMBO J 32: 926-937
- PubMed: 23481256 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/emboj.2013.41
- Primary Citation Related Structures:
4J73, 4J77, 4J78, 4J79, 4J81, 4J82, 4J84, 4J86, 4J87, 4J8B, 4J8G - PubMed Abstract:
Cytoplasmic dilysine motifs on transmembrane proteins are captured by coatomer α-COP and β'-COP subunits and packaged into COPI-coated vesicles for Golgi-to-ER retrieval. Numerous ER/Golgi proteins contain K(x)Kxx motifs, but the rules for their recognition are unclear. We present crystal structures of α-COP and β'-COP bound to a series of naturally occurring retrieval motifs-encompassing KKxx, KxKxx and non-canonical RKxx and viral KxHxx sequences. Binding experiments show that α-COP and β'-COP have generally the same specificity for KKxx and KxKxx, but only β'-COP recognizes the RKxx signal. Dilysine motif recognition involves lysine side-chain interactions with two acidic patches. Surprisingly, however, KKxx and KxKxx motifs bind differently, with their lysine residues transposed at the binding patches. We derive rules for retrieval motif recognition from key structural features: the reversed binding modes, the recognition of the C-terminal carboxylate group which enforces lysine positional context, and the tolerance of the acidic patches for non-lysine residues.
- Memorial Sloan-Kettering Cancer Center, Howard Hughes Medical Institute and the Structural Biology Program, New York, NY, USA.
Organizational Affiliation: