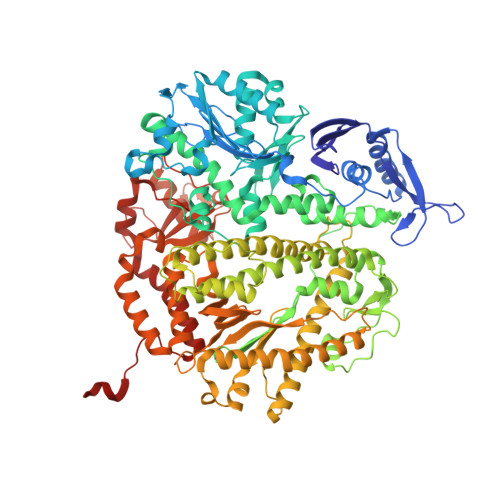
Alteration in the cavity size adjacent to the active site of RB69 DNA polymerase changes its conformational dynamics.
Xia, S., Wood, M., Bradley, M.J., De La Cruz, E.M., Konigsberg, W.H.(2013) Nucleic Acids Res 41: 9077-9089
- PubMed: 23921641 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkt674
- Primary Citation Related Structures:
4J2A, 4J2B, 4J2E - PubMed Abstract:
Internal cavities are a common feature of many proteins, often having profound effects on the dynamics of their interactions with substrate and binding partners. RB69 DNA polymerase (pol) has a hydrophobic cavity right below the nucleotide binding pocket at the tip of highly conserved L415 side chain. Replacement of this residue with Gly or Met in other B family pols resulted in higher mutation rates. When similar substitutions for L415 were introduced into RB69pol, only L415A and L415G had dramatic effects on pre-steady-state kinetic parameters, reducing base selectivity by several hundred fold. On the other hand, the L415M variant behaved like the wild-type. Using a novel tC(o)-tCnitro Förster Resonance Energy Transfer (FRET) assay, we were able to show that the partition of the primer terminus between pol and exonuclease (exo) domains was compromised with the L415A and L415G mutants, but not with the L415M variant. These results could be rationalized by changes in their structures as determined by high resolution X-ray crystallography.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520-8114, USA.
Organizational Affiliation: