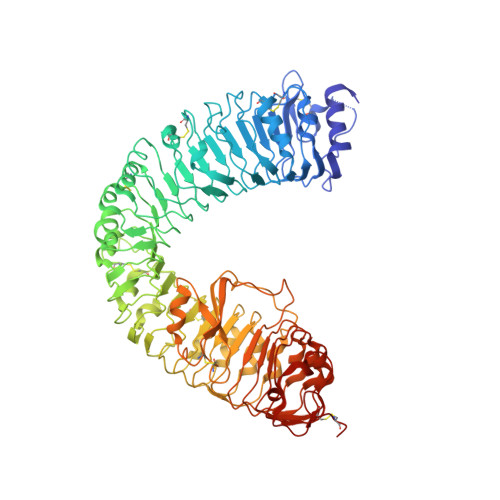
Structural basis for differential recognition of brassinolide by its receptors
She, J., Han, Z., Zhou, B., Chai, J.(2013) Protein Cell 4: 475-482
- PubMed: 23709366 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1007/s13238-013-3027-8
- Primary Citation Related Structures:
4J0M - PubMed Abstract:
Brassinosteroids, a group of plant steroid hormones, regulate many aspects of plant growth and development. We and other have previously solved the crystal structures of BRI1(LRR) in complex with brassinolide, the most active brassinosteroid identified thus far. Although these studies provide a structural basis for the recognition of brassinolide by its receptor BRI1, it still remains poorly understood how the hormone differentiates among its conserved receptors. Here we present the crystal structure of the BRI1 homolog BRL1 in complex with brassinolide. The structure shows that subtle differences around the brassinolide binding site can generate a striking effect on its recognition by the BRI1 family of receptors. Structural comparison of BRL1 and BRI1 in their brassinolide-bound forms reveals the molecular basis for differential binding of brassinolide to its different receptors, which can be used for more efficient design of plant growth regulators for agricultural practice. On the basis of our structural studies and others' data, we also suggest possible mechanisms for the activation of BRI1 family receptors.
- School of Life Sciences, Peking University, Beijing, 100871, China.
Organizational Affiliation: