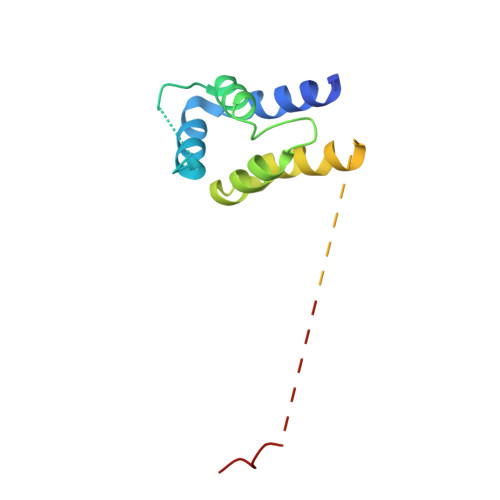
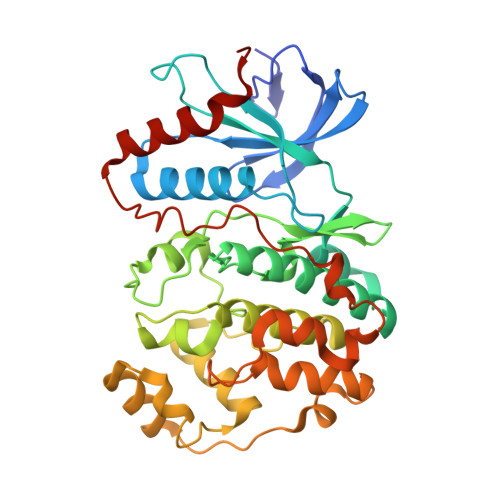
Structure of ERK2 bound to PEA-15 reveals a mechanism for rapid release of activated MAPK.
Mace, P.D., Wallez, Y., Egger, M.F., Dobaczewska, M.K., Robinson, H., Pasquale, E.B., Riedl, S.J.(2013) Nat Commun 4: 1681-1681
- PubMed: 23575685 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms2687
- Primary Citation Related Structures:
4IZ5, 4IZ7, 4IZA - PubMed Abstract:
ERK1/2 kinases are the principal effectors of a central signalling cascade that converts extracellular stimuli into cell proliferation and migration responses and, when deregulated, can promote cell oncogenic transformation. The scaffolding protein PEA-15 is a death effector domain protein that directly interacts with ERK1/2 and affects ERK1/2 subcellular localization and phosphorylation. Here, to understand this ERK1/2 signalling complex, we have solved the crystal structures of PEA-15 bound to three different ERK2 phospho-conformers. The structures reveal that PEA-15 uses a bipartite binding mode, occupying two key docking sites of ERK2. Remarkably, PEA-15 can efficiently bind the ERK2 activation loop in the critical Thr-X-Tyr region in different phosphorylation states. PEA-15 binding triggers an extended allosteric conduit in dually phosphorylated ERK2, disrupting key features of active ERK2. At the same time PEA-15 binding protects ERK2 from dephosphorylation, thus setting the stage for immediate ERK activity upon its release from the PEA-15 inhibitory complex.
- Program in Apoptosis and Cell Death Research, Cancer Center, Sanford-Burnham Medical Research Institute, 10901 North Torrey Pines Road, La Jolla, California 92037, USA.
Organizational Affiliation: