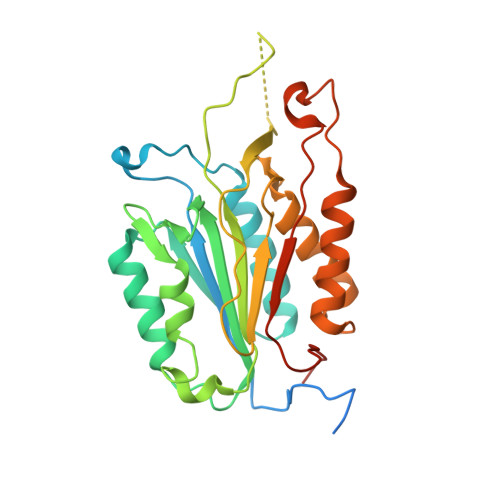
The regulatory mechanism of the caspase 6 pro-domain revealed by crystal structure and biochemical assays.
Cao, Q., Wang, X.J., Li, L.F., Su, X.D.(2014) Acta Crystallogr D Biol Crystallogr 70: 58-67
- PubMed: 24419379 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004713024218
- Primary Citation Related Structures:
4IYR - PubMed Abstract:
Caspase 6 (CASP6) is a neuron degeneration-related protease and is widely considered to be a potential drug-design target against neurodegenerative diseases such as Huntington's disease and Alzheimer's disease. The N-terminal pro-peptide of CASP6, also referred to as the pro-domain, contains 23 residues and its functional role remains elusive. In this study, the crystal structure of a full-length CASP6 zymogen mutant, proCASP6H121A, was solved. Although the pro-domain was flexible in the crystal, without visible electron density, structural analyses combined with biochemical assays revealed that the pro-domain inhibited CASP6 auto-activation by inhibiting intramolecular cleavage at the intersubunit cleavage site TEVD(193) and also by preventing this site from intermolecular cleavage at low protein concentration through a so-called `suicide-protection' mechanism. Further experiments showed that the length of the pro-domain and the side chain of Asn18 played critical roles in suicide protection. These results disclosed a new inhibitory mechanism of CASP6 and shed light on the pathogenesis and therapeutically relevant study of CASP6-related neurodegenerative diseases.
- State Key Laboratory of Protein and Plant Gene Research, and Biodynamic Optical Imaging Center (BIOPIC), School of Life Sciences, Peking University, Beijing 100871, People's Republic of China.
Organizational Affiliation: