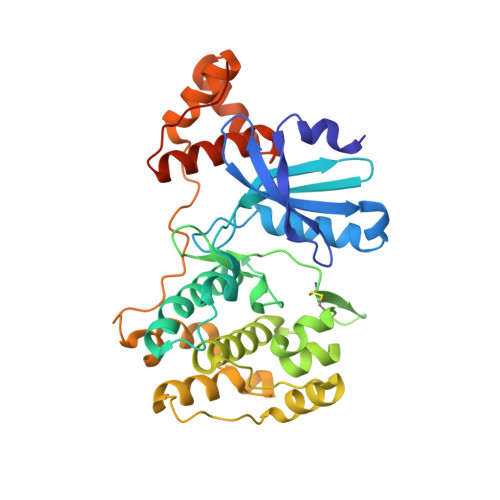
Structural basis for the regulation of maternal embryonic leucine zipper kinase.
Cao, L.S., Wang, J., Chen, Y., Deng, H., Wang, Z.X., Wu, J.W.(2013) PLoS One 8: e70031-e70031
- PubMed: 23922895 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0070031
- Primary Citation Related Structures:
4IXP - PubMed Abstract:
MELK (maternal embryonic leucine zipper kinase), which is a member of the AMPK (AMP-activated protein kinase)-related kinase family, plays important roles in diverse cellular processes and has become a promising drug target for certain cancers. However, the regulatory mechanism of MELK remains elusive. Here, we report the crystal structure of a fragment of human MELK that contains the kinase domain and ubiquitin-associated (UBA) domain. The UBA domain tightly binds to the back of the kinase domain, which may contribute to the proper conformation and activity of the kinase domain. Interestingly, the activation segment in the kinase domain displays a unique conformation that contains an intramolecular disulfide bond. The structural and biochemical analyses unravel the molecular mechanisms for the autophosphorylation/activation of MELK and the dependence of its catalytic activity on reducing agents. Thus, our results may provide the basis for designing specific MELK inhibitors for cancer treatment.
- MOE Key Laboratory of Protein Sciences and Tsinghua-Peking Center for Life Sciences, School of Life Sciences, Tsinghua University, Beijing, China.
Organizational Affiliation: