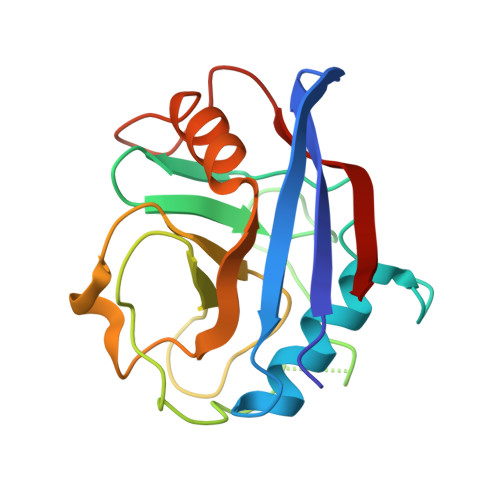
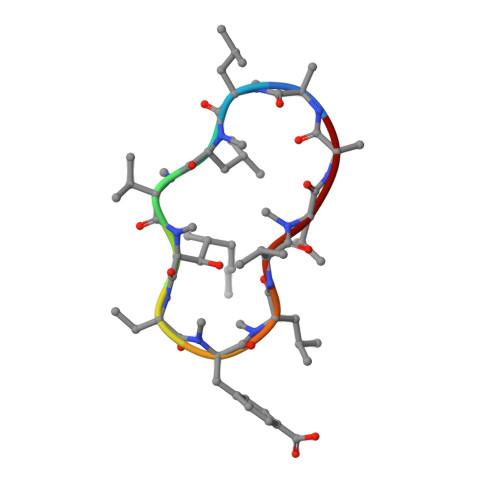
HIV-1 evades innate immune recognition through specific cofactor recruitment.
Rasaiyaah, J., Tan, C.P., Fletcher, A.J., Price, A.J., Blondeau, C., Hilditch, L., Jacques, D.A., Selwood, D.L., James, L.C., Noursadeghi, M., Towers, G.J.(2013) Nature 503: 402-405
- PubMed: 24196705 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature12769
- Primary Citation Related Structures:
4IPZ - PubMed Abstract:
Human immunodeficiency virus (HIV)-1 is able to replicate in primary human macrophages without stimulating innate immunity despite reverse transcription of genomic RNA into double-stranded DNA, an activity that might be expected to trigger innate pattern recognition receptors. We reasoned that if correctly orchestrated HIV-1 uncoating and nuclear entry is important for evasion of innate sensors then manipulation of specific interactions between HIV-1 capsid and host factors that putatively regulate these processes should trigger pattern recognition receptors and stimulate type 1 interferon (IFN) secretion. Here we show that HIV-1 capsid mutants N74D and P90A, which are impaired for interaction with cofactors cleavage and polyadenylation specificity factor subunit 6 (CPSF6) and cyclophilins (Nup358 and CypA), respectively, cannot replicate in primary human monocyte-derived macrophages because they trigger innate sensors leading to nuclear translocation of NF-κB and IRF3, the production of soluble type 1 IFN and induction of an antiviral state. Depletion of CPSF6 with short hairpin RNA expression allows wild-type virus to trigger innate sensors and IFN production. In each case, suppressed replication is rescued by IFN-receptor blockade, demonstrating a role for IFN in restriction. IFN production is dependent on viral reverse transcription but not integration, indicating that a viral reverse transcription product comprises the HIV-1 pathogen-associated molecular pattern. Finally, we show that we can pharmacologically induce wild-type HIV-1 infection to stimulate IFN secretion and an antiviral state using a non-immunosuppressive cyclosporine analogue. We conclude that HIV-1 has evolved to use CPSF6 and cyclophilins to cloak its replication, allowing evasion of innate immune sensors and induction of a cell-autonomous innate immune response in primary human macrophages.
- University College London, Medical Research Council Centre for Medical Molecular Virology, Division of Infection and Immunity, University College London, 90 Gower St, London WC1E 6BT, United Kingdom.
Organizational Affiliation: