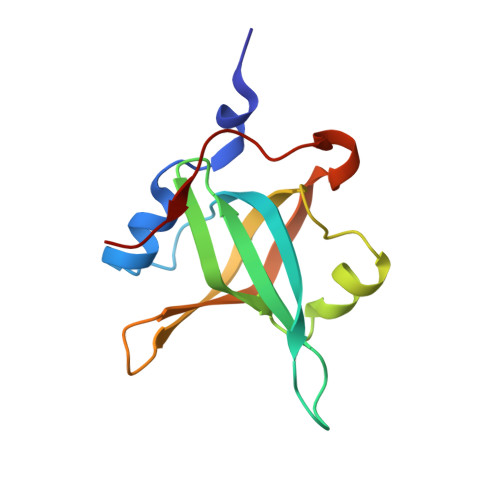
Surface Reengineering of RPA70N Enables Cocrystallization with an Inhibitor of the Replication Protein A Interaction Motif of ATR Interacting Protein.
Feldkamp, M.D., Frank, A.O., Kennedy, J.P., Patrone, J.D., Vangamudi, B., Waterson, A.G., Fesik, S.W., Chazin, W.J.(2013) Biochemistry 52: 6515-6524
- PubMed: 23962067 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi400542z
- Primary Citation Related Structures:
4IPC, 4IPD, 4IPG, 4IPH - PubMed Abstract:
Replication protein A (RPA) is the primary single-stranded DNA (ssDNA) binding protein in eukaryotes. The N-terminal domain of the RPA70 subunit (RPA70N) interacts via a basic cleft with a wide range of DNA processing proteins, including several that regulate DNA damage response and repair. Small molecule inhibitors that disrupt these protein-protein interactions are therefore of interest as chemical probes of these critical DNA processing pathways and as inhibitors to counter the upregulation of DNA damage response and repair associated with treatment of cancer patients with radiation or DNA-damaging agents. Determination of three-dimensional structures of protein-ligand complexes is an important step for elaboration of small molecule inhibitors. However, although crystal structures of free RPA70N and an RPA70N-peptide fusion construct have been reported, RPA70N-inhibitor complexes have been recalcitrant to crystallization. Analysis of the P61 lattice of RPA70N crystals led us to hypothesize that the ligand-binding surface was occluded. Surface reengineering to alter key crystal lattice contacts led to the design of RPA70N E7R, E100R, and E7R/E100R mutants. These mutants crystallized in a P212121 lattice that clearly had significant solvent channels open to the critical basic cleft. Analysis of X-ray crystal structures, target peptide binding affinities, and (15)N-(1)H heteronuclear single-quantum coherence nuclear magnetic resonance spectra showed that the mutations do not result in perturbations of the RPA70N ligand-binding surface. The success of the design was demonstrated by determining the structure of RPA70N E7R soaked with a ligand discovered in a previously reported molecular fragment screen. A fluorescence anisotropy competition binding assay revealed this compound can inhibit the interaction of RPA70N with the peptide binding motif from the DNA damage response protein ATRIP. The implications of the results are discussed in the context of ongoing efforts to design RPA70N inhibitors.
- Department of Biochemistry, ‡Department of Chemistry, §Department of Pharmacology, and ∥Center for Structural Biology, Vanderbilt University , Nashville, Tennessee 37232, United States.
Organizational Affiliation: