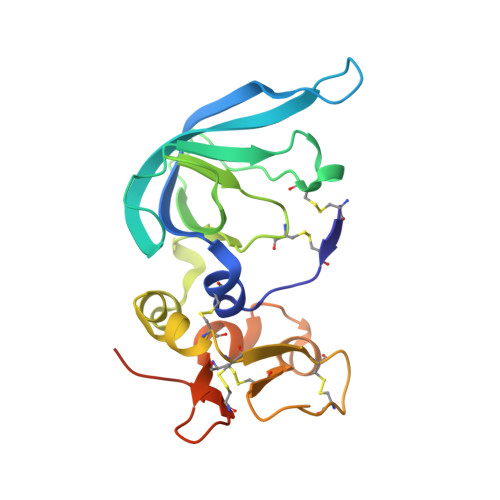
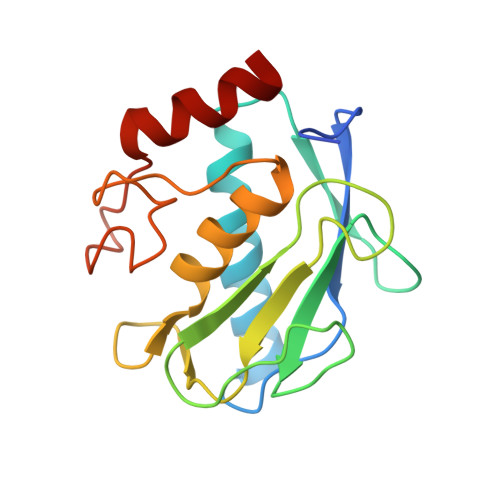
Matrix Metalloproteinase-10/TIMP-2 Structure and Analyses Define Conserved Core Interactions and Diverse Exosite Interactions in MMP/TIMP Complexes.
Batra, J., Soares, A.S., Mehner, C., Radisky, E.S.(2013) PLoS One 8: e75836-e75836
- PubMed: 24073280 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0075836
- Primary Citation Related Structures:
4ILW - PubMed Abstract:
Matrix metalloproteinases (MMPs) play central roles in vertebrate tissue development, remodeling, and repair. The endogenous tissue inhibitors of metalloproteinases (TIMPs) regulate proteolytic activity by binding tightly to the MMP active site. While each of the four TIMPs can inhibit most MMPs, binding data reveal tremendous heterogeneity in affinities of different TIMP/MMP pairs, and the structural features that differentiate stronger from weaker complexes are poorly understood. Here we report the crystal structure of the comparatively weakly bound human MMP-10/TIMP-2 complex at 2.1 Å resolution. Comparison with previously reported structures of MMP-3/TIMP-1, MT1-MMP/TIMP-2, MMP-13/TIMP-2, and MMP-10/TIMP-1 complexes offers insights into the structural basis of binding selectivity. Our analyses identify a group of highly conserved contacts at the heart of MMP/TIMP complexes that define the conserved mechanism of inhibition, as well as a second category of diverse adventitious contacts at the periphery of the interfaces. The AB loop of the TIMP N-terminal domain and the contact loops of the TIMP C-terminal domain form highly variable peripheral contacts that can be considered as separate exosite interactions. In some complexes these exosite contacts are extensive, while in other complexes the AB loop or C-terminal domain contacts are greatly reduced and appear to contribute little to complex stability. Our data suggest that exosite interactions can enhance MMP/TIMP binding, although in the relatively weakly bound MMP-10/TIMP-2 complex they are not well optimized to do so. Formation of highly variable exosite interactions may provide a general mechanism by which TIMPs are fine-tuned for distinct regulatory roles in biology.
- Department of Cancer Biology, Mayo Clinic Cancer Center, Jacksonville, Florida, United States of America.
Organizational Affiliation: