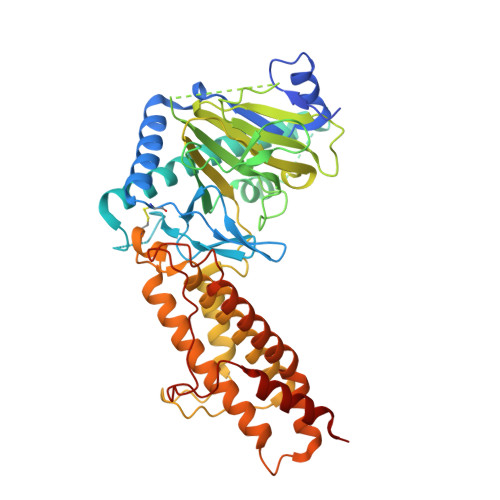
Structural basis for inhibition of the fat mass and obesity associated protein (FTO)
Aik, W.S., Demetriades, M., Hamdan, M.K.K., Bagg, E.A.L., Yeoh, K.K., Lejeune, C., Zhang, Z., McDonough, M.A., Schofield, C.J.(2013) J Med Chem 56: 3680-3688
- PubMed: 23547775 Search on PubMed
- DOI: https://doi.org/10.1021/jm400193d
- Primary Citation Related Structures:
4IDZ, 4IE0, 4IE4, 4IE5, 4IE6, 4IE7 - PubMed Abstract:
The fat mass and obesity associated protein (FTO) is a potential target for anti-obesity medicines. FTO is a 2-oxoglutarate (2OG)-dependent N-methyl nucleic acid demethylase that acts on substrates including 3-methylthymidine, 3-methyluracil, and 6-methyladenine. To identify FTO inhibitors, we screened a set of 2OG analogues and related compounds using differential scanning fluorometry- and liquid chromatography-based assays. The results revealed sets of both cyclic and acyclic 2OG analogues that are FTO inhibitors. Identified inhibitors include small molecules that have been used in clinical studies for the inhibition of other 2OG oxygenases. Crystallographic analyses reveal inhibition by 2OG cosubstrate or primary substrate competitors as well as compounds that bind across both cosubstrate and primary substrate binding sites. The results will aid the development of more potent and selective FTO inhibitors.
- Chemistry Research Laboratory, University of Oxford , 12 Mansfield Road, Oxford OX1 3TA, United Kingdom.
Organizational Affiliation: