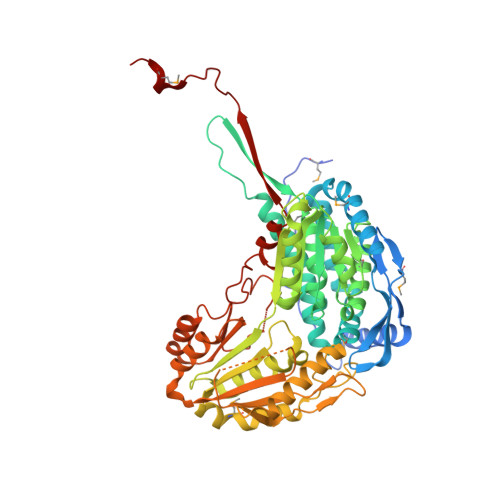
Characterization of the proline-utilization pathway in Mycobacterium tuberculosis through structural and functional studies.
Lagautriere, T., Bashiri, G., Paterson, N.G., Berney, M., Cook, G.M., Baker, E.N.(2014) Acta Crystallogr D Biol Crystallogr 70: 968-980
- PubMed: 24699642
- DOI: https://doi.org/10.1107/S1399004713034391
- Primary Citation Related Structures:
4IDM, 4IDS, 4IHI, 4JDC - PubMed Abstract:
The proline-utilization pathway in Mycobacterium tuberculosis (Mtb) has recently been identified as an important factor in Mtb persistence in vivo, suggesting that this pathway could be a valuable therapeutic target against tuberculosis (TB). In Mtb, two distinct enzymes perform the conversion of proline into glutamate: the first step is the oxidation of proline into Δ(1)-pyrroline-5-carboxylic acid (P5C) by the flavoenzyme proline dehydrogenase (PruB), and the second reaction involves converting the tautomeric form of P5C (glutamate-γ-semialdehyde) into glutamate using the NAD(+)-dependent Δ(1)-pyrroline-5-carboxylic dehydrogenase (PruA). Here, the three-dimensional structures of Mtb-PruA, determined by X-ray crystallography, in the apo state and in complex with NAD(+) are described at 2.5 and 2.1 Å resolution, respectively. The structure reveals a conserved NAD(+)-binding mode, common to other related enzymes. Species-specific conformational differences in the active site, however, linked to changes in the dimer interface, suggest possibilities for selective inhibition of Mtb-PruA despite its reasonably high sequence identity to other PruA enzymes. Using recombinant PruA and PruB, the proline-utilization pathway in Mtb has also been reconstituted in vitro. Functional validation using a novel NMR approach has demonstrated that the PruA and PruB enzymes are together sufficient to convert proline to glutamate, the first such demonstration for monofunctional proline-utilization enzymes.
- Structural Biology Laboratory, School of Biological Sciences and Maurice Wilkins Centre for Molecular Biodiscovery, University of Auckland, Auckland 1010, New Zealand.
Organizational Affiliation: