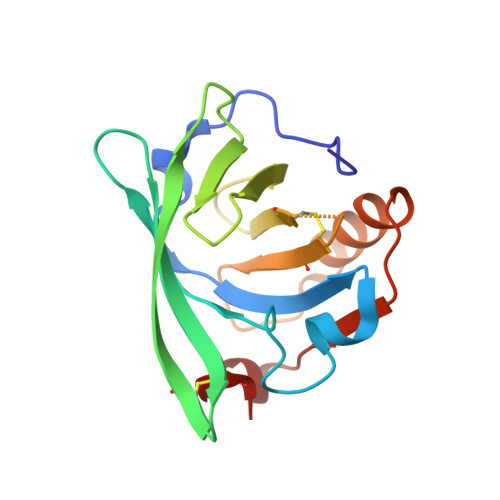
The differences in binding 12-carbon aliphatic ligands by bovine beta-lactoglobulin isoform A and B studied by isothermal titration calorimetry and X-ray crystallography
Loch, J.I., Bonarek, P., Polit, A., Swiatek, S., Dziedzicka-Wasylewska, M., Lewinski, K.(2013) J Mol Recognit 26: 357-367
- PubMed: 23784992 Search on PubMed
- DOI: https://doi.org/10.1002/jmr.2280
- Primary Citation Related Structures:
4IB6, 4IB7, 4IB8, 4IB9, 4IBA - PubMed Abstract:
Isoforms A (LGB-A) and B (LGB-B) of bovine lactoglobulin, the milk protein, differ in positions 64 (D↔G) and 118 (V↔A). Interactions of LGB-A and LGB-B with sodium dodecyl sulfate (SDS), dodecyltrimethylammonium chloride (DTAC) and lauric acid (LA), 12-carbon ligands possessing differently charged polar groups, were investigated using isothermal titration calorimetry and X-ray crystallography, to study the proton linkage phenomenon and to distinguish between effects related to different isoforms and different ligand properties. The determined values of ΔS and ΔH revealed that for all ligands, binding is entropically driven. The contribution from enthalpy change is lower and shows strong dependence on type of buffer that indicates proton release from the protein varying with protein isoform and ligand type and involvement of LA and Asp64 (in isoform A) in this process. The ligand affinities for both isoforms were arranged in the same order, DTAC < LA < SDS, and were systematically lower for variant B. The entropy change of the complexation process was always higher for isoform A, but these values were compensated by changes in enthalpy, resulting in almost identical ΔG for complexes of both isoforms. The determined crystal structures showed that substitution in positions 64 and 118 did not influence the overall structure of LGB complexes. The chemical character of the ligand polar group did not affect the position of its aliphatic chain in protein β-barrel, indicating a major role of hydrophobic interactions in ligand binding that prevailed even with the repulsion between positively charged DTAC and lysine residues located at binding site entrance.
- Jagiellonian University, Faculty of Chemistry, Department of Crystal Chemistry and Crystal Physics, Ingardena 3, 30-060 Kraków, Poland.
Organizational Affiliation: