An asymmetric SMC-kleisin bridge in prokaryotic condensin
Burmann, F., Shin, H.C., Basquin, J., Soh, Y.M., Gimenez-Oya, V., Kim, Y.G., Oh, B.H., Gruber, S.(2013) Nat Struct Mol Biol 20: 371-379
- PubMed: 23353789 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2488
- Primary Citation Related Structures:
3ZGX, 4I98, 4I99 - PubMed Abstract:
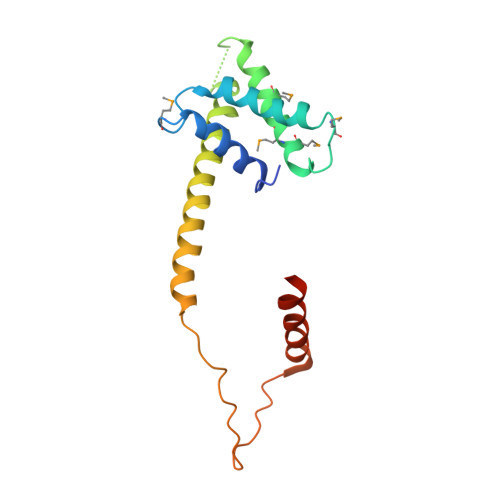
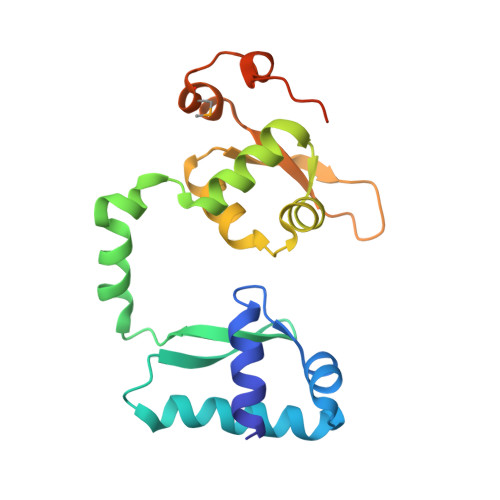
Eukaryotic structural maintenance of chromosomes (SMC)-kleisin complexes form large, ring-shaped assemblies that promote accurate chromosome segregation. Their asymmetric structural core comprises SMC heterodimers that associate with both ends of a kleisin subunit. However, prokaryotic condensin Smc-ScpAB is composed of symmetric Smc homodimers associated with the kleisin ScpA in a postulated symmetrical manner. Here, we demonstrate that Smc molecules have two distinct binding sites for ScpA. The N terminus of ScpA binds the Smc coiled coil, whereas the C terminus binds the Smc ATPase domain. We show that in Bacillus subtilis cells, an Smc dimer is bridged by a single ScpAB to generate asymmetric tripartite rings analogous to eukaryotic SMC complexes. We define a molecular mechanism that ensures asymmetric assembly, and we conclude that the basic architecture of SMC-kleisin rings evolved before the emergence of eukaryotes.
- Max Planck Research Group Chromosome Organization and Dynamics, Max Planck Institute of Biochemistry, Martinsried, Germany.
Organizational Affiliation: