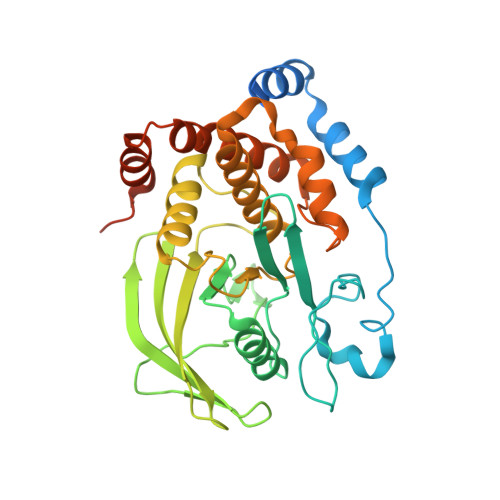
X-Ray Structure of PTP1B in Complex with a New PTP1B Inhibitor.
Reddy, M.V., Ghadiyaram, C., Panigrahi, S.K., Krishnamurthy, N.R., Hosahalli, S., Chandrasekharappa, A.P., Manna, D., Badiger, S.E., Dubey, P.K., Mangamoori, L.N.(2014) Protein Pept Lett 21: 90-93
- PubMed: 23964742 Search on PubMed
- DOI: https://doi.org/10.2174/09298665113209990089
- Primary Citation Related Structures:
4I8N - PubMed Abstract:
Protein tyrosine phosphatase 1B (PTP1B) is a prototype non receptor cytoplasmic PTPase enzyme that has been implicated in regulation of insulin and leptin signaling pathways. Studies on PTP1B knockout mice and PTP1B antisense treated mice suggested that inhibition of PTP1B would be an effective strategy for the treatment of type II diabetes and obesity. Here we report the X-ray structure of PTP1B in complex with compound IN1834-146C (PDB ID 4I8N). The crystals belong to P3121 space group with cell dimensions (a = b = 87.89 Å, c = 103.68 Å) diffracted to 2.5 Å. The crystal structure contained one molecule of protein in the asymmetric unit and was solved by molecular replacement method. The compound engages both catalytic site and allosteric sites of PTP1B protein. We described the molecular interaction of the compound with the active site residues of PTP1B in this crystal structure report.
- Aurigene Discovery Technologies Ltd, 39-40 KIADB Industrial Area, Phase II, Electronic city, Hosur Road, Bangalore-560100, India. manusekhar1975@gmail.com.
Organizational Affiliation: