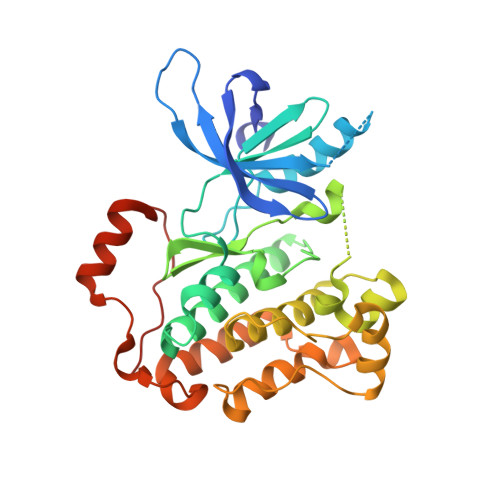
Insights into the Aberrant Activity of Mutant EGFR Kinase Domain and Drug Recognition.
Gajiwala, K.S., Feng, J., Ferre, R., Ryan, K., Brodsky, O., Weinrich, S., Kath, J.C., Stewart, A.(2013) Structure 21: 209-219
- PubMed: 23273428 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2012.11.014
- Primary Citation Related Structures:
4I1Z, 4I20, 4I21, 4I22, 4I23, 4I24 - PubMed Abstract:
The oncogenicity of the L858R mutant form of the epidermal growth factor receptor (EGFR) in non-small-cell lung cancer is thought to be due to the constitutive activation of its kinase domain. The selectivity of the marketed drugs gefitinib and erlotinib for L858R mutant is attributed to their specific recognition of the active kinase and to weaker ATP binding by L858R EGFR. We present crystal structures showing that neither L858R nor the drug-resistant L858R+T790M EGFR kinase domain is in the constitutively active conformation. Additional co-crystal structures show that gefitinib and dacomitinib, an irreversible anilinoquinazoline derivative currently in clinical development, may not be conformation specific for the active state of the enzyme. Structural data further reveal the potential mode of recognition of one of the autophosphorylation sites in the C-terminal tail, Tyr-1016, by the kinase domain. Biochemical and biophysical evidence suggest that the oncogenic mutations impact the conformational dynamics of the enzyme.
- Cancer Structural Biology, Oncology Medicinal Chemistry, Pfizer Worldwide Research and Development, 10770 Science Center Drive, San Diego, CA 92121, USA. ketan.gajiwala@pfizer.com
Organizational Affiliation: