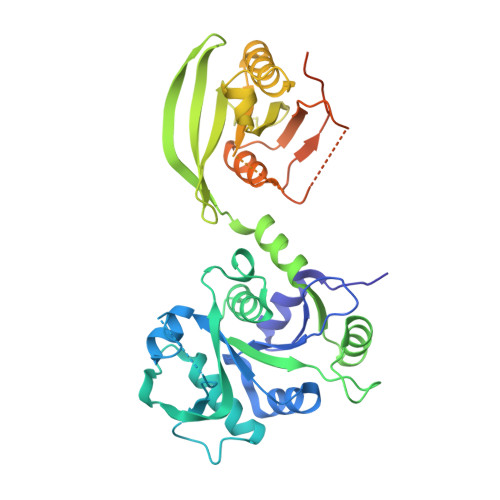
The crystal structure reveals the molecular mechanism of bifunctional 3,4-dihydroxy-2-butanone 4-phosphate synthase/GTP cyclohydrolase II (Rv1415) from Mycobacterium tuberculosis
Singh, M., Kumar, P., Yadav, S., Gautam, R., Sharma, N., Karthikeyan, S.(2013) Acta Crystallogr D Biol Crystallogr 69: 1633-1644
- PubMed: 23999287 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444913011402
- Primary Citation Related Structures:
4I14 - PubMed Abstract:
The enzymes 3,4-dihydroxy-2-butanone 4-phosphate synthase (DHBPS) and GTP cyclohydrolase II (GCHII) catalyze the initial steps of both branches of the bacterial riboflavin-biosynthesis pathway. The structures and molecular mechanisms of DHBPS and GCHII as separate polypeptides are known; however, their organization and molecular mechanism as a bifunctional enzyme are unknown to date. Here, the crystal structure of an essential bifunctional DHBPS/GCHII enzyme from Mycobacterium tuberculosis (Mtb-ribA2) is reported at 3.0 Å resolution. The crystal structure revealed two conformationally different molecules of Mtb-ribA2 in the asymmetric unit that form a dimer via their GCHII domains. Interestingly, analysis of the crystal packing revealed a long `helical-like oligomer' formed by DHBPS and GCHII functional homodimers, thus generating an `open-ended' unit-cell lattice. However, size-exclusion chromatography studies suggest that Mtb-ribA2 exists as a dimer in solution. To understand the discrepancy between the oligomerization observed in solution and in the crystal structure, the DHBPS (Mtb-DHBPS) and GCHII (Mtb-GCHII) domains of Mtb-ribA2 have been cloned, expressed and purified as His-tagged proteins. Size-exclusion chromatography studies indicated that Mtb-GCHII is a dimer while Mtb-DHBPS exists as a monomer in solution. Moreover, kinetic studies revealed that the GCHII activities of Mtb-ribA2 and Mtb-GCHII are similar, while the DHBPS activity of Mtb-ribA2 is much higher than that of Mtb-DHBPS alone. Taken together, the results strongly suggest that Mtb-ribA2 exists as a dimer formed through its GCHII domains and requires full-length Mtb-ribA2 for optimal DHBPS activity.
- CSIR - Institute of Microbial Technology, Council of Scientific and Industrial Research (CSIR), Sector 39-A, Chandigarh 160 036, India.
Organizational Affiliation: