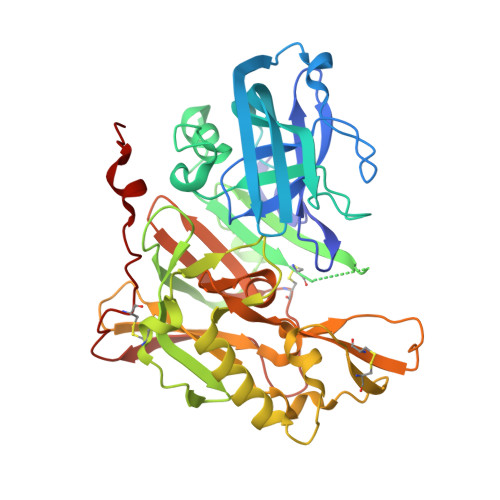
Structure-based design of novel dihydroisoquinoline BACE-1 inhibitors that do not engage the catalytic aspartates.
Bowers, S., Xu, Y.Z., Yuan, S., Probst, G.D., Hom, R.K., Chan, W., Konradi, A.W., Sham, H.L., Zhu, Y.L., Beroza, P., Pan, H., Brecht, E., Yao, N., Lougheed, J., Tam, D., Ren, Z., Ruslim, L., Bova, M.P., Artis, D.R.(2013) Bioorg Med Chem Lett 23: 2181-2186
- PubMed: 23465612
- DOI: https://doi.org/10.1016/j.bmcl.2013.01.103
- Primary Citation of Related Structures:
4HZT, 4I0G, 4I0Z, 4I10, 4I11 - PubMed Abstract:
The structure-activity relationship of a series of dihydroisoquinoline BACE-1 inhibitors is described. Application of structure-based design to screening hit 1 yielded sub-micromolar inhibitors. Replacement of the carboxylic acid of 1 was guided by X-ray crystallography, which allowed the replacement of a key water-mediated hydrogen bond. This work culminated in compounds such as 31, which possess good BACE-1 potency, excellent permeability and a low P-gp efflux ratio.
- Department of Chemical Sciences, Elan Pharmaceuticals, 180 Oyster Point Boulevard, South San Francisco, CA 94080, USA. simeongbowers@gmail.com
Organizational Affiliation: