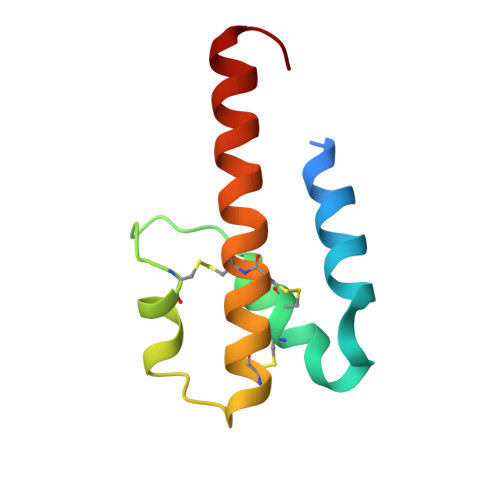
Human C3a and C3a desArg anaphylatoxins have conserved structures, in contrast to C5a and C5a desArg.
Bajic, G., Yatime, L., Klos, A., Andersen, G.R.(2013) Protein Sci 22: 204-212
- PubMed: 23184394 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2200
- Primary Citation Related Structures:
4HW5, 4HWJ - PubMed Abstract:
Complement is a part of innate immunity that has a critical role in the protection against microbial infections, bridges the innate with the adaptive immunity and initiates inflammation. Activation of the complement, by specific recognition of molecular patterns presented by an activator, for example, a pathogen cell, in the classical and lectin pathways or spontaneously in the alternative pathway, leads to the opsonization of the activator and the production of pro-inflammatory molecules such as the C3a anaphylatoxin. The biological function of this anaphylatoxin is regulated by carboxypeptidase B, a plasma protease that cleaves off the C-terminal arginine yielding C3a desArg, an inactive form. While functional assays demonstrate strikingly different physiological effects between C3a and C3a desArg, no structural information is available on the possible conformational differences between the two proteins. Here, we report a novel and simple expression and purification protocol for recombinant human C3a and C3a desArg anaphylatoxins, as well as their crystal structures at 2.3 and 2.6 Å, respectively. Structural analysis revealed no significant conformational differences between the two anaphylatoxins in contrast to what has been reported for C5a and C5a desArg. We compare the structures of different anaphylatoxins and discuss the relevance of their observed conformations to complement activation and binding of the anaphylatoxins to their cognate receptors.
- Department of Molecular Biology and Genetics, Aarhus University, Gustav Wieds Vej 10C, DK-8000 Aarhus, Denmark.
Organizational Affiliation: