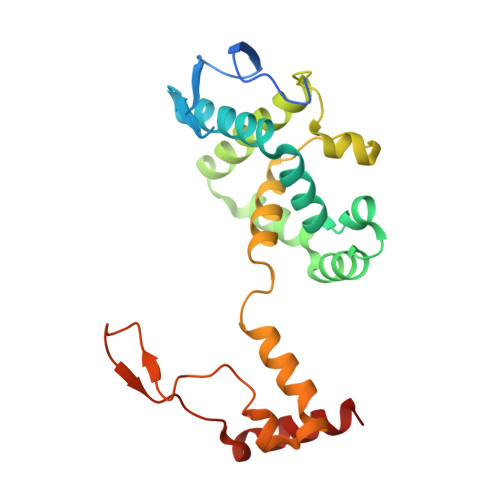
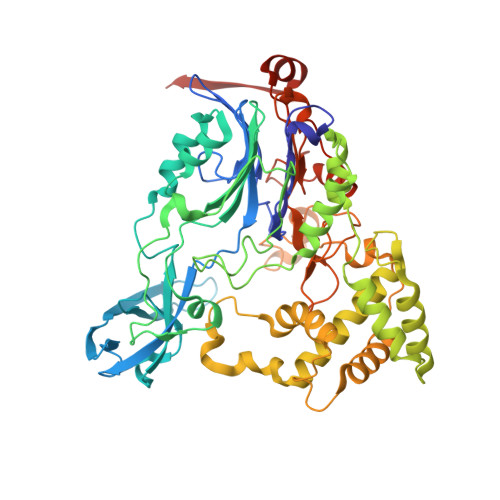
Structure of a class III engineered cephalosporin acylase: comparisons with class I acylase and implications for differences in substrate specificity and catalytic activity.
Golden, E., Paterson, R., Tie, W.J., Anandan, A., Flematti, G., Molla, G., Rosini, E., Pollegioni, L., Vrielink, A.(2013) Biochem J 451: 217-226
- PubMed: 23373797 Search on PubMed
- DOI: https://doi.org/10.1042/BJ20121715
- Primary Citation Related Structures:
4HSR, 4HST - PubMed Abstract:
The crystal structure of the wild-type form of glutaryl-7-ACA (7-aminocephalosporanic acid) acylase from Pseudomonas N176 and a double mutant of the protein (H57βS/H70βS) that displays enhanced catalytic efficiency on cephalosporin C over glutaryl-7-aminocephalosporanic acid has been determined. The structures show a heterodimer made up of an α-chain (229 residues) and a β-chain (543 residues) with a deep cavity, which constitutes the active site. Comparison of the wild-type and mutant structures provides insights into the molecular reasons for the observed enhanced specificity on cephalosporin C over glutaryl-7-aminocephalosporanic acid and offers the basis to evolve a further improved enzyme variant. The nucleophilic catalytic serine residue, Ser(1β), is situated at the base of the active site cavity. The electron density reveals a ligand covalently bound to the catalytic serine residue, such that a tetrahedral adduct is formed. This is proposed to mimic the transition state of the enzyme for both the maturation step and the catalysis of the substrates. A view of the transition state configuration of the enzyme provides important insights into the mechanism of substrate binding and catalysis.
- School of Chemistry and Biochemistry, University of Western Australia, 35 Stirling Highway, Crawley, WA 6009, Australia.
Organizational Affiliation: